Mohapatra Research Group
Contact
Dr. Lishibanya Mohapatra
Assistant Professor
College of Science
School of Physics and Astronomy
lxmsps@rit.edu
The Mohapatra Group is looking for students. We are hiring!
Central Question
Just like human bodies have organs to complete various biological functions, living cells have distinguishable subcellular parts called organelles, each specifically designed for its own specialized task. These organelles have been observed since the dawn of microscopy, but only recently have we developed experimental tools to begin to investigate how they attain their characteristic shapes and sizes, and how that relates to their specific functions. Using theory and simulations, and in collaboration with several experimental groups, the Mohapatra group studies the key principles required to assemble structures inside cells - a world smaller than the width of a human hair.
About Lishibanya Mohapatra
 Lishibanya Mohapatra, Ph.D. is an Assistant Professor in the School of Physics and Astronomy at the Rochester Institute of Technology. Currently, her research is focused on understanding how living cells measure and control the size of their organelles. Dr. Mohapatra is the Vice-chair of RIT's Women in Science (WISe) program and a Faculty Research Mentor of the HHMI-funded Inclusive Excellence program at RIT. She enjoys working with students on a variety of research problems at the interface of Physics and Biology.
Lishibanya Mohapatra, Ph.D. is an Assistant Professor in the School of Physics and Astronomy at the Rochester Institute of Technology. Currently, her research is focused on understanding how living cells measure and control the size of their organelles. Dr. Mohapatra is the Vice-chair of RIT's Women in Science (WISe) program and a Faculty Research Mentor of the HHMI-funded Inclusive Excellence program at RIT. She enjoys working with students on a variety of research problems at the interface of Physics and Biology.
Call for Students
Mohapatra group is always looking for motivated students to work with on a variety of research projects. Curiosity and open communication are important prerequisites, along with a desire or skill to code. If interested, please contact Dr. Lishi via her email.
Undergraduate students can participate in projects that range from code optimization and analyzing experimental images and data to process development. They will be provided opportunities to communicate results to the general community through peer-reviewed publications and oral and poster presentations.
Graduate and PhD students will lead a research project related to organelle assembly, building mathematical models and using experimental data to limit the parameter space of model variables. They will communicate results to scientists through peer reviewed publications and oral and poster presentations.
Post-doctoral associate will conduct analytical and computational research on the mechanisms of organelle assembly, mediated by related cellular proteins. They will create hypothesis-driven mechanistic mathematical models, with an emphasis on making predictions that can be experimentally tested. They will analyze experimental data, compare theory results with experimental data, and communicate results to scientists through peer reviewed publications and oral and poster presentations.
Current Projects
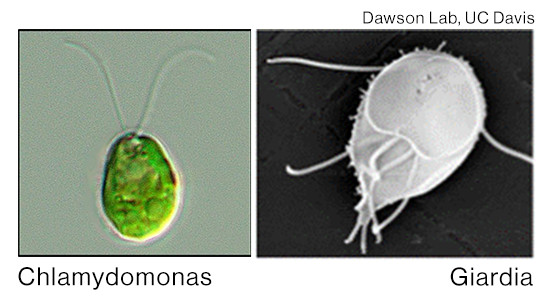
All living cells contain organelles which serve specific functions related to their characteristic sizes and shapes. These organelles grow in a shared pool of building blocks, and interestingly, cohabitating organelles can achieve a variety of configurations in a shared resource environment. For example, two flagella of Chlamydomonas are identical in size, while four pairs of flagella in Giardia maintain different sizes. How do they do it? We investigate the various puzzles that arise while cohabitating structures by assembled by living cells.
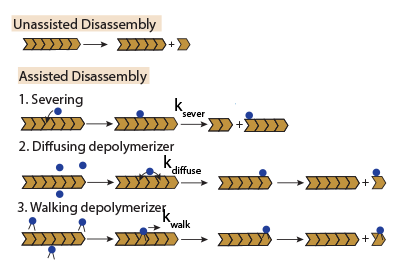
Structures within cells are assembled and disassembled rapidly while exchanging their building block proteins with those in the pool in the order of seconds. To achieve this rapid exchange, cells use several disassembly proteins to remove subunits from cytoskeletal structures using distinct molecular mechanisms. We use stochastic simulations to quantitatively study the specific action of these disassembly proteins, individually and as a collection.
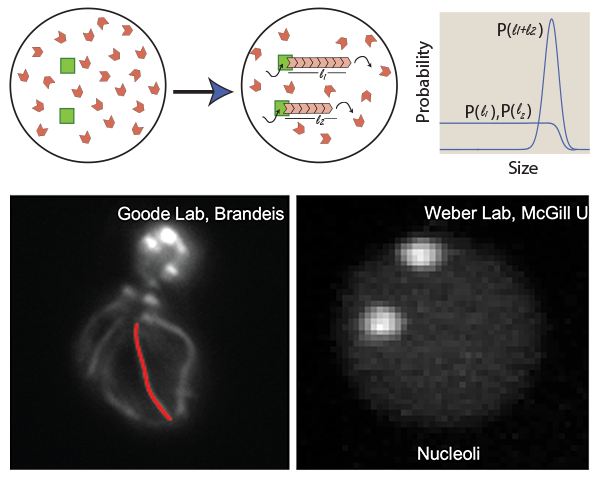
In cells, we see several examples of organelles which maintain their characteristic size while sharing a common pool of building blocks. While linear actin cables and spherical patches share a common pool of actin monomers in yeast cells, C. elegans forms two equal-sized spherical nucleoli in the nucleus every cell cycle while sharing a common pool of nucleolar proteins. We use mathematical modeling and simulations to study the strategies used by cells to share resources among multiple organelles in the same pool.
Mohapatra Research Group News
Mohapatra Group welcomes capstone students Ashley Lasko, Ben Russell, and research assistant Rivka Stasavage!
Dr. Lishi is awarded the Maximizing Investigators' Research Award (MIRA) from the National Institute of General Medical Sciences (NIH) to build a 5-year research program to study organelle assembly.
Mohapatra group presented four posters at the RIT Undergraduate Research Symposium.
Mohapatra Group presented a series of four short talks at the RIT soft matter meeting.
Mohapatra group welcomes four new students in the summer of 2022 – Owen Cheevers, Mimi Harrison, Huayuan(Harry) Han, and Stanley Liang.
Michaela Cohen completed her Capstone project with flying colors – congrats!
Mohapatra Group welcomes Capstone student Michaela Cohen!
Pete Miller is awarded the Emerson award to conduct research in the Mohapatra Group over the summer of 2021.
Dr. Lishi is awarded the SRS seed funding award to study the assembly of nucleolus.
Dr. Lishi is awarded the Dean’s Research Initiation grant to study the size regulation of nucleolus.
Publications
National Institute of Health
Amount: $ 1.77 million
Active Collaborations

My philosophy on scientific collaboration:
I place myself firmly in the camp of interdisciplinary collaborations and cooperation. I enjoy working with people with different expertise, focus, and interests. Students are considered equal collaborators in my group and open communication and creativity are strongly encouraged.

Owen Cheevers
Third-year (physics)

Mimi Harrison
Second-year (physics) and Inclusive Excellence fellow.
Outside class, she enjoys reading, astronomy, and spending time outside.

Pete Miller, fifth-year (applied mathematics/physics) double major

Michaela Cohen, fifth-year (BS/MS physics)
Huayuan(Harry) Han, Fourth-year (physics)
Stanley Liang, Fourth-year Physics Major

Chloe Shiff, Graduate Student, Stanford University
Rabeya Hussaini, Graduate student, NYU
- Thomas Fai, Brandeis University
- Jane Kondev, Brandeis University
- Shashank Sekhar, Emory University
- Steph Weber, McGill University
Other Interests
Dr. Lishi is the Vice - Chair of Women in Science and Engineering (WISe) at RIT.
"As a member of WISe at RIT, I aim to support women faculty and students by organizing and participating in outreach events to inspire future generations of women scientists."
Dr. Lishi is an Inclusive Excellence Faculty member.
"As an IE faculty Research Mentor, I participate in workshops focused on mentoring students from historically underrepresented groups with a goal to improve student experience in my research group."
Dr. Lishi was a Science Communication Fellow and member of the Brandeis SciComm Lab 2018-2020.
"As Communication Fellow, I offered individual coaching, organized targeted workshops, and a range of initiatives to support the scientific community with the skills they need to communicate their scientific endeavors to any audience in written, spoken or visual form."

Dr. Lishi was the Co-Director of the Quantitative Biology Research Community (QBReC) at Brandeis University from 2016-2020.
"QBReC is an HHMI-funded program for Brandeis undergraduate students, which gives them a unique opportunity to experience an academic environment by facilitating research opportunities with Brandeis faculty who are working at the interface of the physical and life sciences. Research projects were designed and led by student advisors, and I mentored both the advisors and undergraduate students throughout the process. In collaboration with the Experiential Learning Group at Brandeis, I developed team-building activities focused on cultivating a community of like-minded scholars."
Dr. Lishi has been a Research Assistant in the summer courses of Physiology and Physical Biology of the Cell at the Marine Biological Laboratory in Wood's Hole, MA.
"I advised biology graduate and post-doctoral students on how to incorporate modeling and programming in their research to tackle problems in biology using MATLAB."
Banner Photo Credits (L to R):
Spindle: O'Connell and Khodjakov, J Cell Sci 2007
Chlamydomonas: Morga and Bastin, Cilia 2013
Budding yeast: Image courtesy of Goode Lab (Brandeis University)
Nucleolus: Image Courtesy of Weber Lab (McGill University)
Microvilli: Crawley et al., JCB 2014




