Lea Michel Research Group

Michel Research Group
Congratulations Louis, Pani, and Abby on your publication now available in the journal Biology Open!
I have been an ASBMB member for 16 years, during which time I have had the opportunity to attend ASBMB conferences, judge poster competitions, write for ASBMB Today, and serve as Chair of the Maximizing Access Committee. I am deeply grateful for the opportunities ASBMB has provided, and I am honored to have been selected as a 2026 ASBMB Fellow.
So proud of Grace Jones for publishing an article in ASBMB TODAY on her research and personal experience with crystallins and cataracts!
The Michel Research Group celebrated the end of the semester and wished Mya Wynn and Grace Perna congratulations as they leave RIT to embark on new adventures in graduate school!

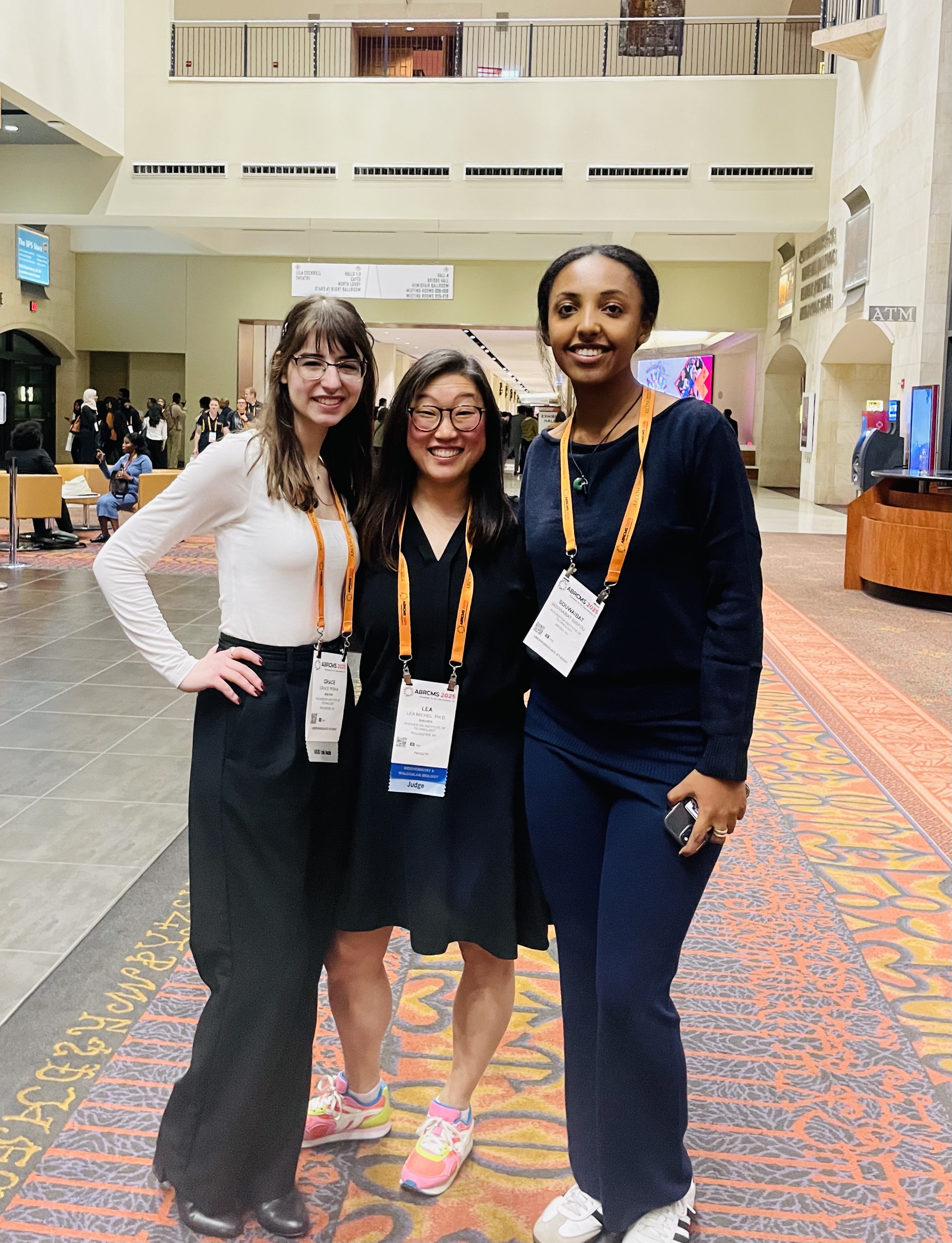
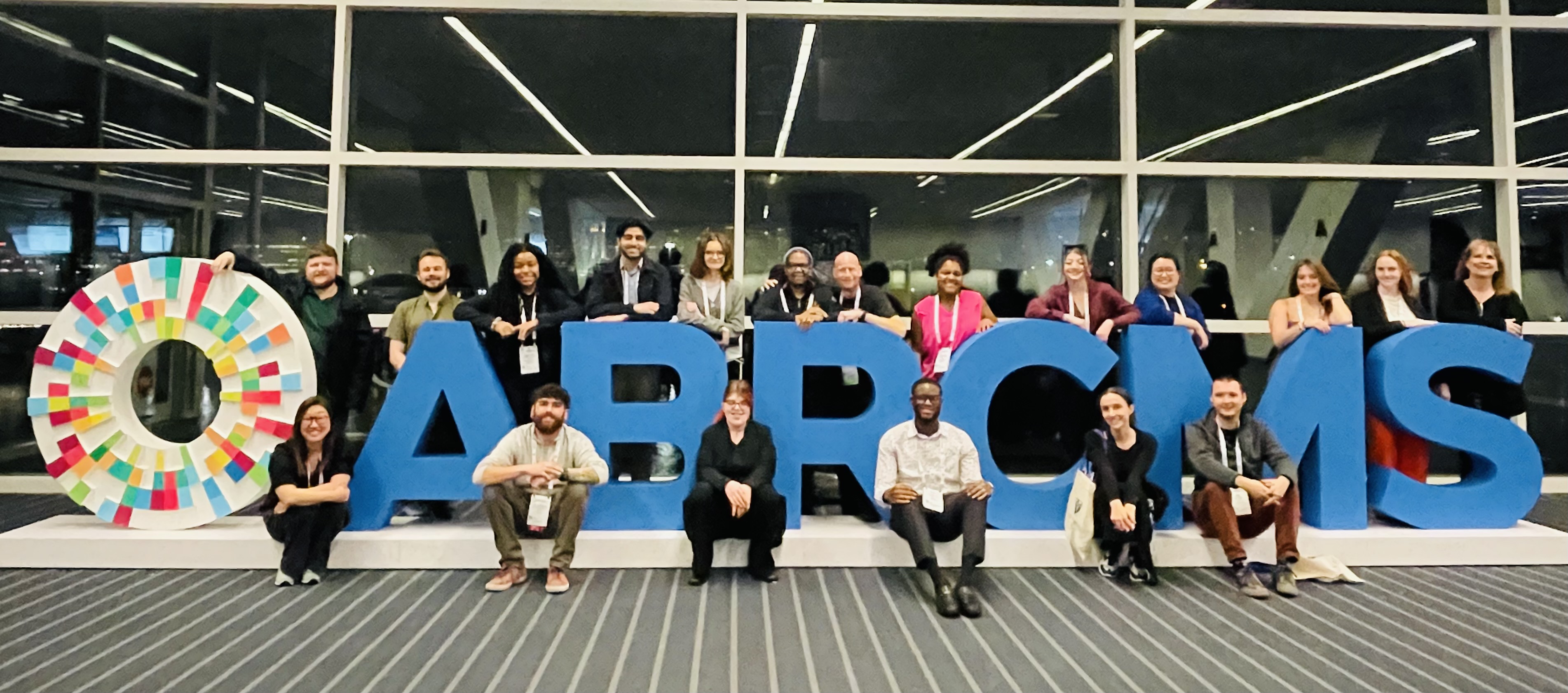
Congratulations Grace and Halley on presenting posters at ABRCMS in San Antonio (November 2025)!

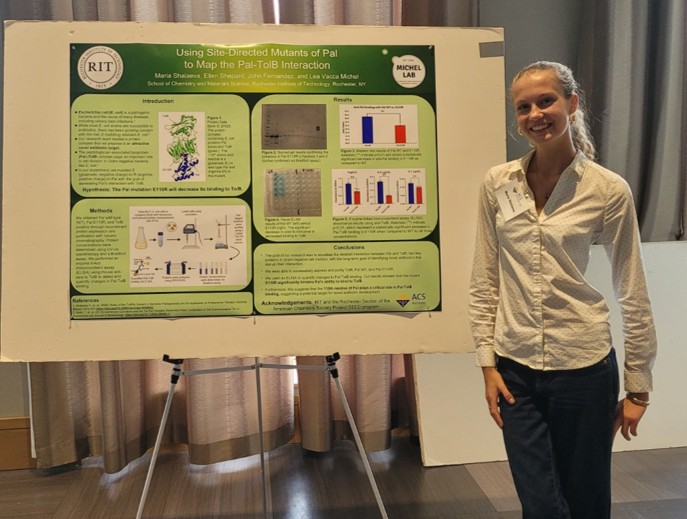
A huge congratulations to our Summer 2025 ACS Project SEED Intern, Maria, who presented her research and won Best Poster Award at the Rochester ACS Sci-Tech Symposium!
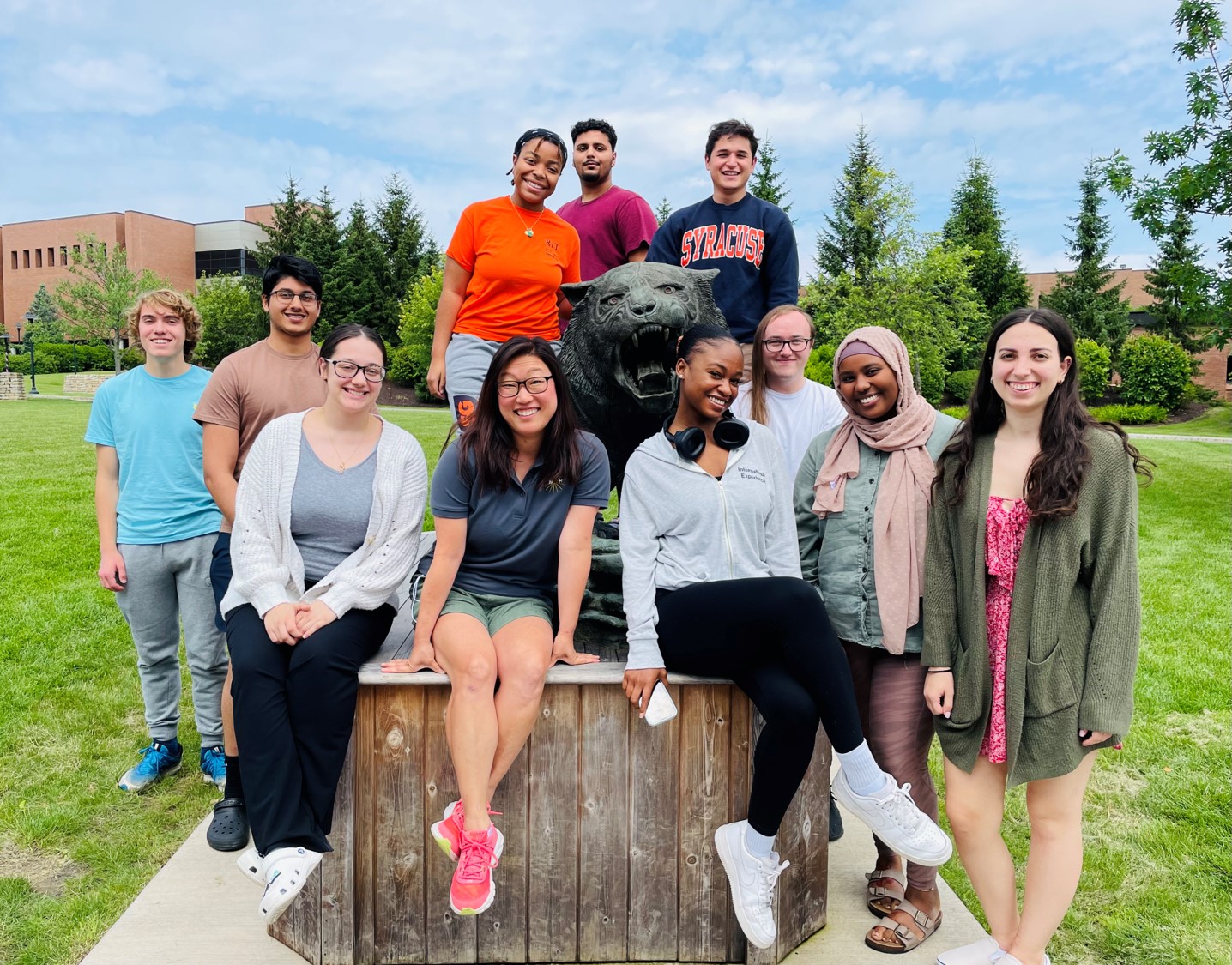
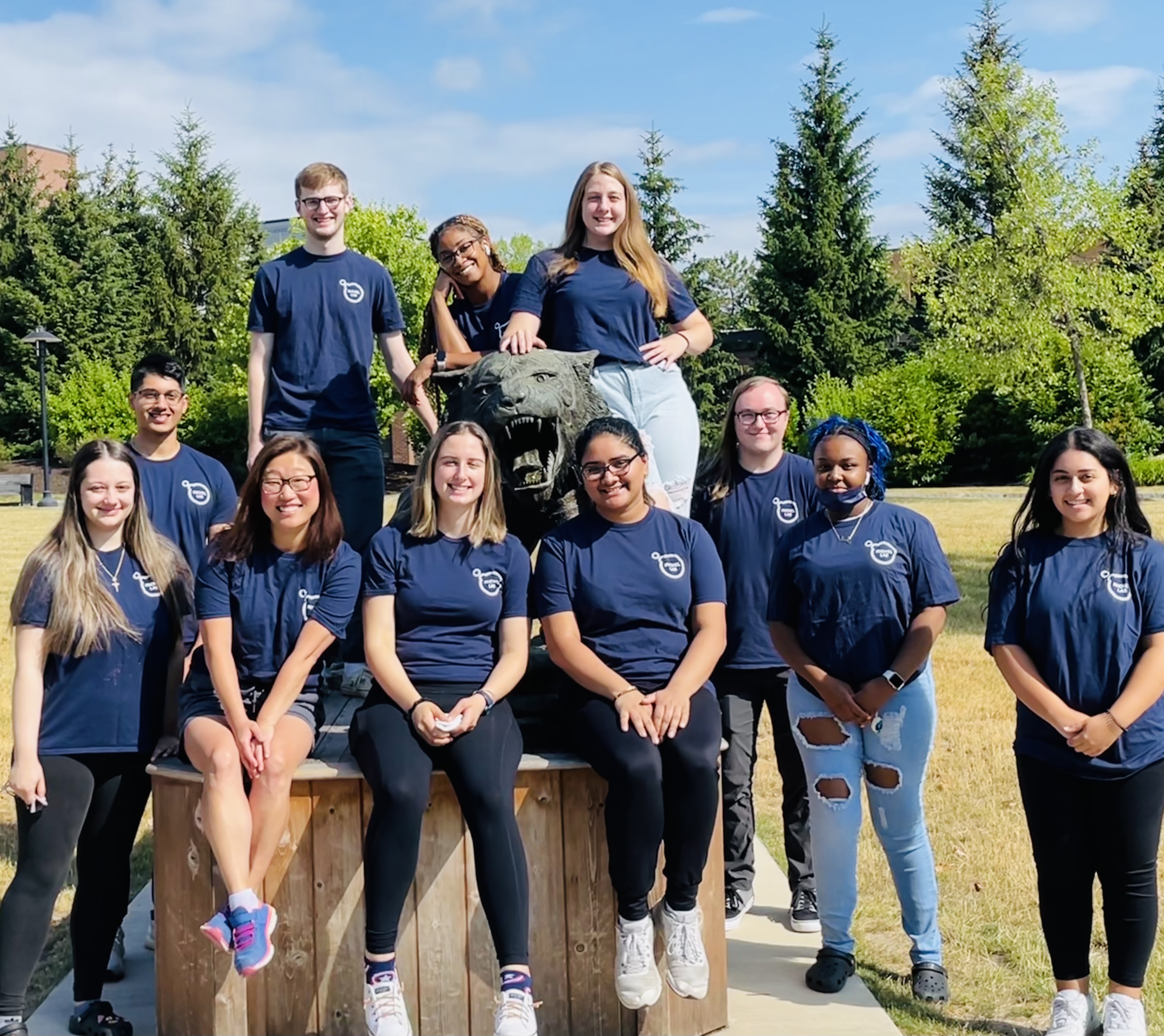
Summer 2025 in the Michel Lab!
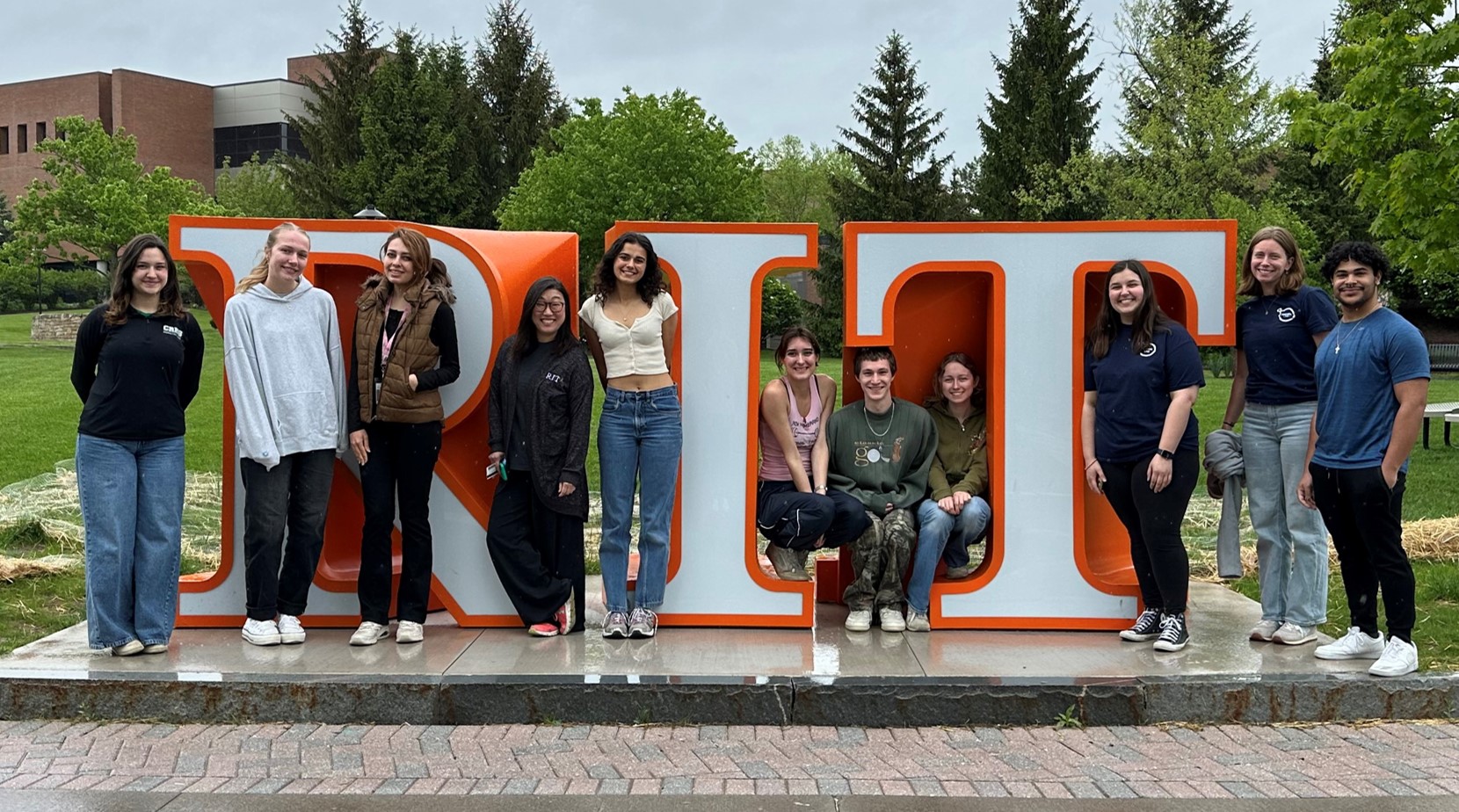

Congratulations, Mia, Guerline, Christina, Aidan, and Grace! So proud of everything you've done and excited about the amazing things you will accomplish in your future!

In April 2025, Mia, Christina, Mya, Nikita, Grace, and Halley presented their research at the national ASBMB conference in Chicago. What an amazing opportunity for these burgeoning scientists!

It was such an honor to receive the 2024-2025 Eisenhart Award for Outstanding Teaching, which is RIT’s highest honor for tenured faculty.


CONGRATULATIONS to Grace Perna (left) for receiving a highly competitive and prestigious 2025 Barry Goldwater Scholarship!

Check out this great article on research opportunities, which was included in the RIT President's Report, featuring our very own Halley Deme!
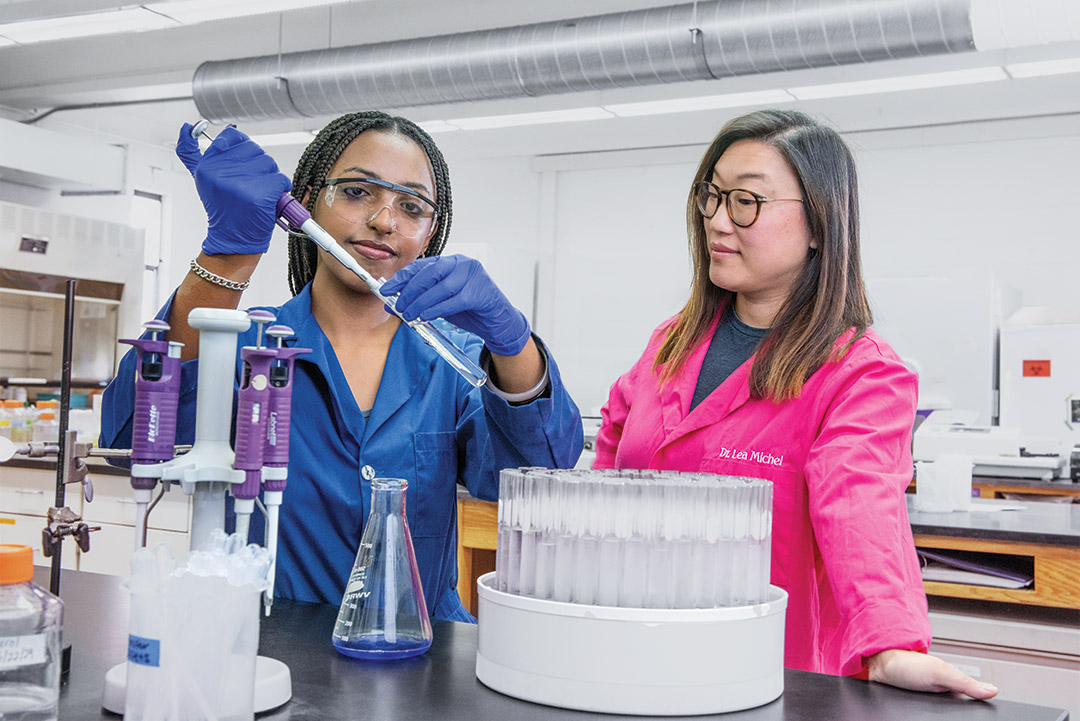

Congratulations to Pani Torabian on her first publication as an RIT Biomedical Engineering PhD Candidate! There are also 8 former Michel Lab undergraduate research students who are coauthors on the paper, along with our collaborators, Dr. Thomas Gaborski and Dr. Anthony Pietropaoli.
Torabian P, Singh N, Crawford J, Gonzalez G, Burgado N, Videva M, Miller A, Perdue J, Dinu M, Pietropaoli A, Gaborski T, Michel LV (2025) The effect of clinically relevant antibiotics on bacterial extracellular vesicle release from E. coli. Int. J Antimicrob Agents 65: 107384.

Michel Lab Holiday Party 2024- We made science snow globes!

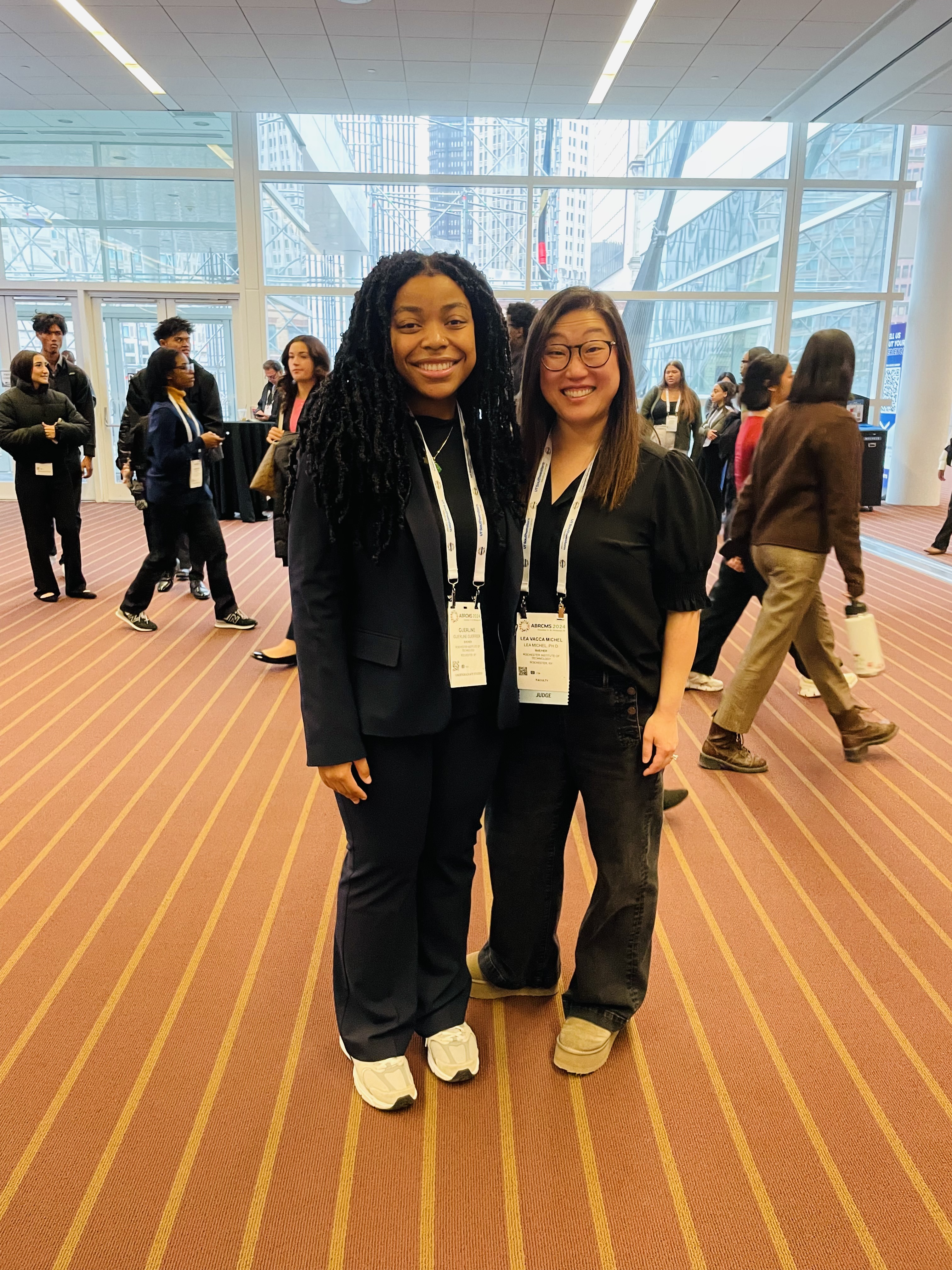
So proud of Guerline and all the RIT folks who presented at ABRCMS 2024 in Pittsburgh!

Since joining the faculty at NTID, Dr. Pepsi Holmquist (Michel research group alumna) has been collaborating with the Michel group on several research projects, including a new NSF project focused on creating signs for Biochemistry. Pepsi and Lea were recently invited by the American Society for Microbiology to write an article on enhancing accessibility for deaf and hard-of-hearing scientists- Check it out!

https://asm.org/Articles/2024/September/Building-Accessibility-for-Deaf-and-Hard-of-Hearin
Overview
 The Michel Research Group was established in 2009. The Principal Investigator, Dr. Lea Vacca Michel, is a Professor in the School of Chemistry and Materials Science at RIT. Dr. Michel is a biophysicist by training and currently works in the fields of protein biochemistry and structural biology. Since 2009, the Michel Group has had over 100 research students (undergraduate and MS). Dr. Michel recruits and inspires a diverse group of research students, and she works hard to create an inclusive research environment where everyone feels welcomed and valued.
The Michel Research Group was established in 2009. The Principal Investigator, Dr. Lea Vacca Michel, is a Professor in the School of Chemistry and Materials Science at RIT. Dr. Michel is a biophysicist by training and currently works in the fields of protein biochemistry and structural biology. Since 2009, the Michel Group has had over 100 research students (undergraduate and MS). Dr. Michel recruits and inspires a diverse group of research students, and she works hard to create an inclusive research environment where everyone feels welcomed and valued.
About Dr. Michel
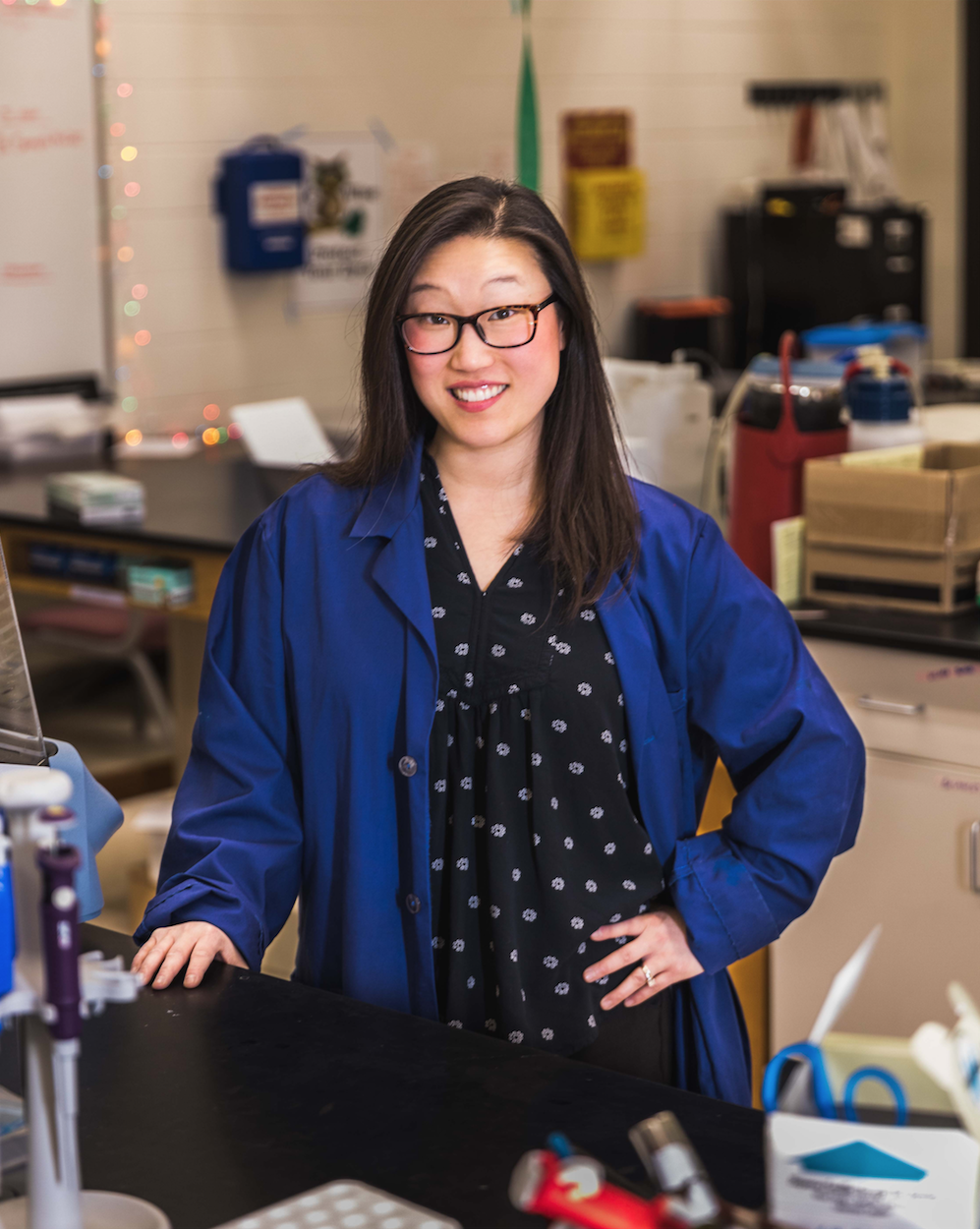

Lea Vacca Michel, Ph.D. is a Professor in the School of Chemistry and Materials Science at the Rochester Institute of Technology and the College of Science's Director of Access and Belonging. Currently, her research is focused on the role of proteins in disease. Dr. Michel is a proud member of RIT's Women in Science (WISe) program (former Chair of WISe), a mentor for the Rochester Project SEED program (former Director), member of the Rochester ACS Women Chemists Committee (WCC), former Director of the Research Strand for the HHMI-funded Inclusive Excellence program at RIT, and current Chair of ASBMB's Maximizing Access Committee. She strives to increase the participation of women and underrepresented minorities (including those who are deaf and hard-of-hearing) in science and math. Dr. Michel has been featured in articles in Nature (Nature 558, 149-151, 2018) and Chemistry World magazine (Chemistry World, Careers), and she was awarded the 2022 ASBMB Early-Career Leadership Award (ASBMB Today article) and the 2023 ChemCUR Outstanding Mentor Award from the Chemistry Division and the Council on Undergraduate Research. In 2024, Dr. Michel was named a Finalist for the Presidential Awards for Excellence in Science, Mathematics, and Engineering Mentoring.
Education and Positions
- B.A. in Physics and Math (Magna Cum Laude), Colgate University, 2002
- M.S. and Ph.D. in Biophysics, University of Rochester, 2007
- Adjunct Professor, SUNY Brockport and Colgate University, Spring 2007
- Postdoctoral Researcher, Cornell University, 2007-2008
- Contract Scientist, Kelly Services (Ortho-Clinical Diagnostics), 2008-2009
- Assistant Professor, School of Chemistry and Materials Science, RIT, 2009-2015
- Associate Professor, School of Chemistry and Materials Science, RIT, 2015-2023
- Director of Access and Belonging for the College of Science, RIT, 2022-present
- Full Professor, School of Chemistry and Materials Science, RIT, 2023-present
Learn more about Dr. Lea Vacca Michel at her official RIT directory page.
Research
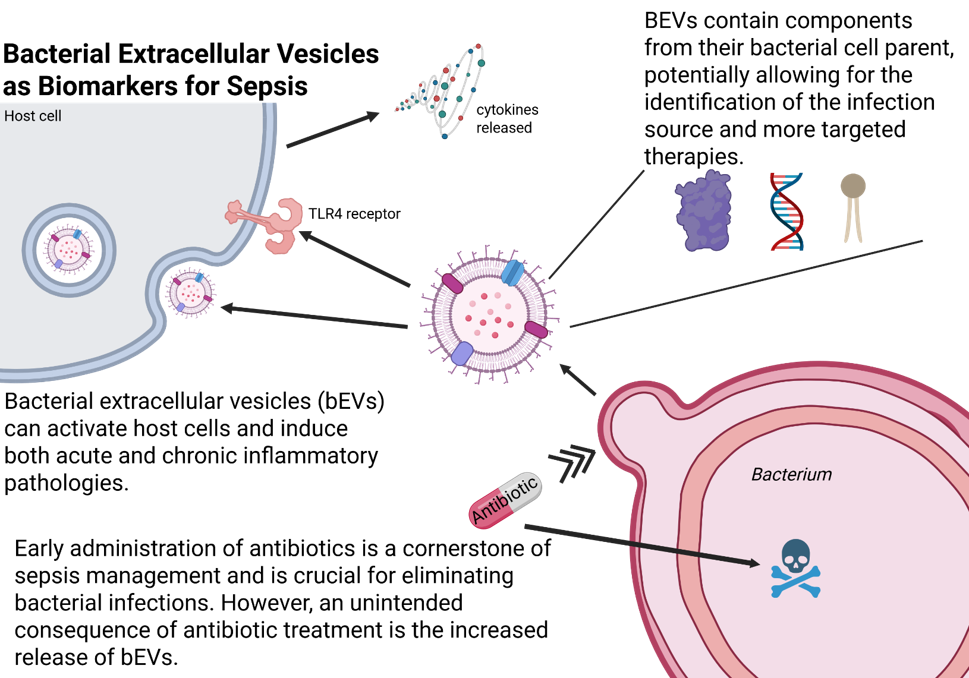
Sepsis is not only the most expensive condition treated in US hospitals, costing close to $37 million/year, but also a leading cause of death, with 350,000 sepsis-related deaths each year. Sepsis occurs when host proinflammatory immune responses become abnormally elevated due to a dysregulated host-response to infection, most commonly caused by bacteria.
The Michel team and collaborators hypothesize that bacterial extracellular vesicles (bEVs) would make attractive diagnostic biomarkers for Gram-negative infection and sepsis. Extracellular vesicles are membrane-enclosed nanobodies that are naturally released from all cell types. Bacterial EVs are generated by both Gram-positive and Gram-negative bacterial cells, containing a multitude of cellular components (protein, DNA, etc.) that originate from the parent bacterium from which they derive.
BEVs are thought to play important roles in pathogenesis, bacterial survival under stress conditions, and interbacterial transport and communication. Because bEVs contain inflammatory molecules, such as LPS, virulence factors, adhesins, and lipoproteins, they are thought to play key roles in endothelial activation and inflammation during the transition from infection to sepsis. BEVs are also capable of self-entry deep into host tissues, resulting in long-term, chronic responses and inflammatory pathologies.
What makes bEVs such attractive diagnostic biomarkers for sepsis: A) bEVs contain unique components from their bacterial cell parent, allowing for the identification of the infection source and more targeted therapies; B) bEVs can activate host cells and induce both acute and chronic inflammatory, sepsis-related pathologies C) bEVs are robust - unlike their bacterial cell “parent,” bEVs can withstand the inundation of broad-spectrum antibiotics.
Michel LV and Gaborski T (2022) Invited Review: Outer Membrane Vesicles as Molecular Biomarkers for Gram-negative Sepsis: Taking Advantage of Nature’s Perfect Packages. J Biol Chem 298(10) 102483. doi: 10.1016/j.jbc.2022.102483
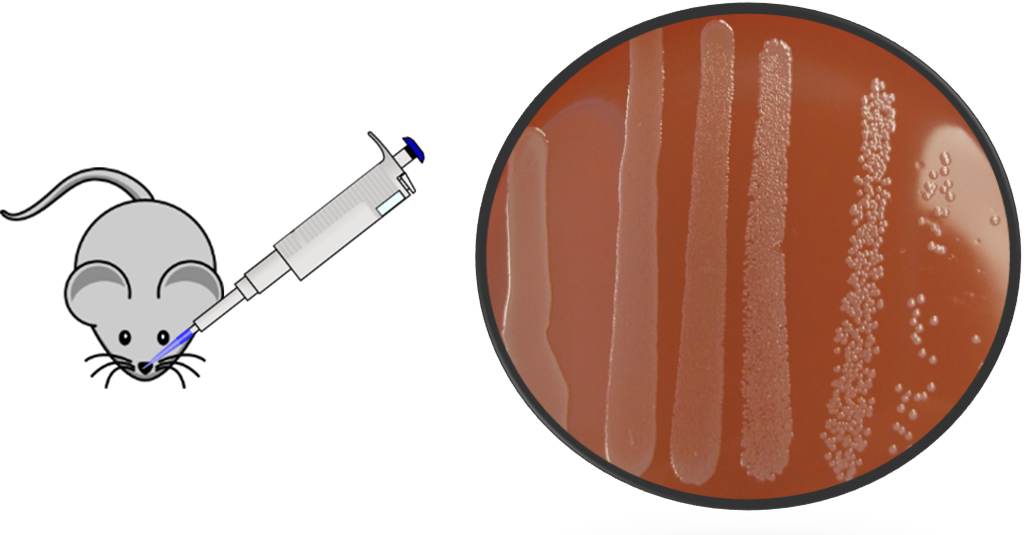
Nontypeable Haemophilus influenzae (NTHi) is a commensal in the human nasopharynx and the cause of pneumonia, meningitis, sinusitis, acute exacerbations of chronic obstructive pulmonary disease, and acute otitis media (AOM). AOM (ear infection) is the most common ailment for which antibiotics are prescribed in the US. With the emergence of new strains of antibiotic resistant bacteria, finding an effective and broad coverage vaccine to protect against AOM-causing pathogens has become a priority. The Michel Research Group employs a biochemical approach toward evaluating conserved outer membrane proteins from NTHi as vaccine candidates. The team has recently begun work characterizing and assessing the effectiveness of a genetically engineered chimeric lipoprotein vaccine, using bacterial extracellular vesicles as a vaccine delivery platform.
The Michel Research Group also developed an NTHi AOM model in C57BL/6J mice to test vaccine candidates. This work is part of a long-standing collaboration between the Michel Research Group and the research group of Dr. Michael Pichichero, Director of the Research Institute at Rochester General Hospital.

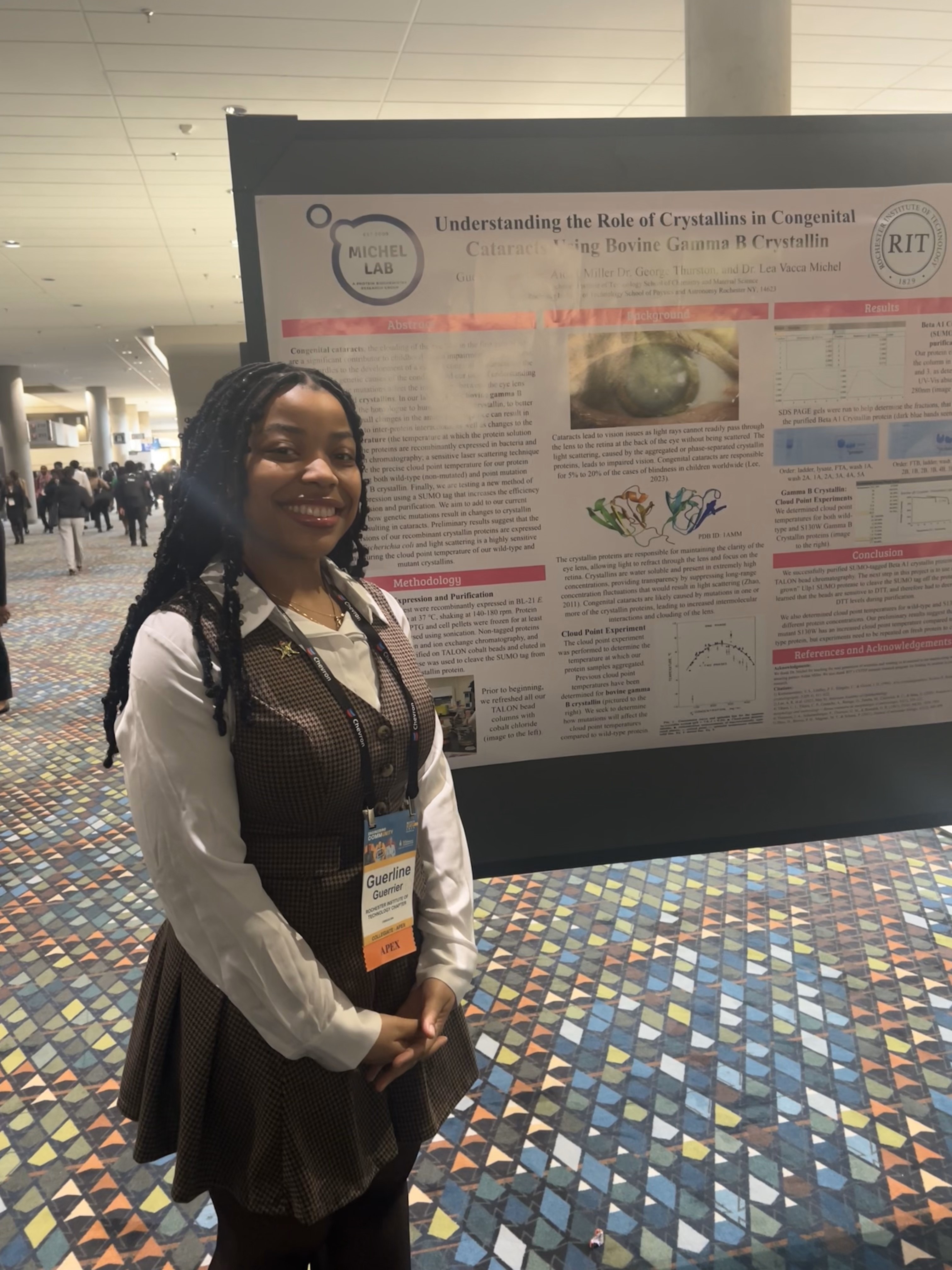
Gamma B crystallin is the bovine homolog to the human lens gamma D crystallin protein. Abnormalities, such as single point mutations, in crystallin proteins have been implicated in cataracts, a condition where loss of protein solubility in the eye lens leads to partial or total blindness due to lens cloudiness. The Michel Research Group collaborates with the Thurston Research Group (PI of the project: Dr. George Thurston, Physics) to better understand how and under what conditions the gamma B crystallins interact with each other, as well as how point mutations within the gamma B crystallin sequence affect those interactions. The collaborative team uses biophysical methods, such as NMR spectroscopy, light scattering, small angle x-ray scattering, and computational modeling, to elucidate the complex interprotein interactions between the crystallins.
Publications
Publications:
*RIT Students
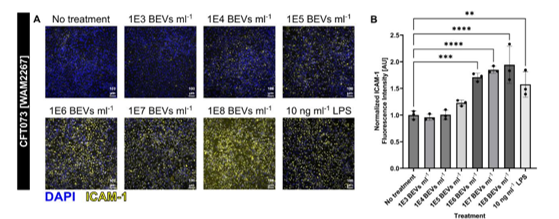
Widom LP*, Torabian P*, Wojehowski AC*, Ghaemmaghami S, Michel LV, Gaborski TR (2026) Antibiotic treatment modulates Escherichia coli-derived bacterial extracellular vesicle (BEV) production and their capacity to upregulate ICAM-1 in human endothelial cells. Biology Open.doi: 10.1242/bio.062476
Torabian P*, Singh N*, Crawford J*, Gonzalez G*, Burgado N*, Videva M*, Miller A*, Perdue J*, Dinu M*, Pietropaoli A, Gaborski T, Michel LV. (2025) The effect of clinically relevant antibiotics on bacterial extracellular vesicle release from E. coli. Int. J Antimicrob Agents 65: 107384. doi: 10.1016/j.ijantimicag.2024.107384
Michel LV, Kaur R, Gleghorn ML, Holmquist M, Pryharski K, Perdue J*, Jones SP*, Jackson N*, Pilo I*, Kasper A*, Labbe N*, Pichichero ME (2022) Haemophilus influenzae Protein D Antibody Suppression in a Multi-component vaccine formulation. FEBS Open Bio. doi: 10.1002/2211-5463.13498
Jones SP*, Cook KH*, Holmquist ML, Almekinder L*, DeLaney A*, Charles R*, Labbe N*, Perdue J*, Jackson N*, Pichichero ME, Kaur R, Michel LV, Gleghorn ML (2022) Vaccine target and carrier molecule nontypeable Haemophilus influenzae Protein D dimerizes like the close Escherichia coli GlpQ homolog but unlike other known homolog dimers. Proteins. doi: 10.1002/prot.26418
Michel LV and Gaborski T (2022) Invited Review: Outer Membrane Vesicles as Molecular Biomarkers for Gram-negative Sepsis: Taking Advantage of Nature’s Perfect Packages. J Biol Chem 298(10) 102483. doi: 10.1016/j.jbc.2022.102483
Wyatt B, Magalhaes RM, Newman D, Michel LV, Newman D (2021) Development of a Faculty Learning Community to Foster Inclusive Research Mentoring, Journal of Faculty Development 35 (2): 44-49.
Gehret A, Trussell J, Michel LV (2021) Experiential Education of Deaf and Hard of Hearing Students in the Lab with Non-Signing Advisors, International Journal of Inclusive Education. https://doi.org/10.1080/13603116.2021.1879948
Michel LV, *Gallardo L, Konovalova A, *Bauer M, *Jackson N, *Zavorin M, *McNamara C, *Pierce J, *Cheng S, *Snyder E, Hellman J, Pichichero ME (2020) Ampicillin triggers the release of Pal in toxic vesicles from Escherichia coli, International Journal of Antimicrobial Agents. doi: 10.1016/j.ijantimicag.2020.106163
Michel LV, Kaur R, *Zavorin M, Pryharski K, Khan MN, *LaClair C, *O’Neil M, Xu Q, Pichichero ME (2018) Intranasal coinfection model allows for assessment of protein vaccines against nontypeable Haemophilus influenzae in mice, Journal of Medical Microbiology 67: 1527-1532. doi: 10.1099/jmm.0.000827
*Sgheiza V, *Novick B, *Stanton S, *Pierce J, *Kalmeta B, *Holmquist MF, *Grimaldi K, Bren KL, Michel LV (2017) Covalent bonding of heme to protein prevents heme capture by nontypeable Haemophilus influenzae, FEBS Open Bio 7: 1778-1783. doi: 10.1002/2211-5463.12324
Gehret AU, Trussell JW, Michel LV (2017) Approaching Undergraduate Research with Students who are Deaf and Hard-of-Hearing, Journal of Science Education for Students with Disabilities 20 (1): Article 4. doi: 10.14448/jsesd.08.0002
Michel LV, *Shaw J, *MacPherson V, *Barnard D, *Bettinger J, *D’Arcy B, Surendran N, Hellman J, Pichichero ME (2015) Dual orientation of the outer membrane lipoprotein Pal in Escherichia coli, Microbiology 161: 1251-1259. doi: 10.1099/mic.0.000084
Michel LV, *Snyder J, *Schmidt R, *Milillo J, *Grimaldi K, *Kalmeta B, Khan N, Sharma S, Wright LK, Pichichero ME (2013) Dual orientation of the outer membrane lipoprotein P6 of nontypeable Haemophilus influenzae, J Bacteriology 195: 3252-3259. doi: 10.1128/JB.00185-13
Craig PA, Michel LV, Bateman RC (2013) A Survey of Educational Uses of Molecular Visualization Freeware, Biochemistry and Molecular Biology Education 41: 193-205. doi: 10.1002/bmb.20693
Peterson JE, Zurakowski D, Italiano JE, Michel LV, Connors S, Oenick M, D’Amato RJ, Klement GL, Folkman MJ (2012) VEGF, PF4 and PDGF are elevated in platelets of colorectal cancer patients, Angiogenesis 15: 265-273. doi: 10.1007/s10456-012-9259-z
Michel LV, *Kalmeta B, *McCreary M, *Snyder J, Craig P, Pichichero ME (2011) Vaccine candidate P6 of nontypable Haemophilus influenzae is not a transmembrane protein based on protein structural analysis, Vaccine 29: 1624-1627. doi: 10.1016/j.vaccine.2010.12.082
Chang A, Kaur R, Michel LV, Casey JR, Pichichero ME (2011) Haemophilus influenzae vaccine candidate outer membrane protein P6 is not conserved in all strains, Hum Vaccines 7: 102-105. doi: 10.4161/hv.7.1.13351 Peterson JE, Zurakowski D, Italiano JE, Michel LV, Fox L, Klement GL, Folkman J (2010) Normal ranges of angiogenesis regulatory proteins in human platelets, Am J Hematology 85: 487-493. doi: 10.1002/ajh.21732
Michel LV, Bren KL (2008) Submolecular unfolding units of Pseudomonas aeruginosa cytochrome c551, J Biol Inorg Chem 13: 837-845. doi: 10.1007/s00775-008-0370-y
Ye T, Kaur R, Senguin FT, Michel LV, Bren KL, Elliott SJ (2008) Methionine ligand lability of type I cytochromes c: Detection of ligand loss using protein film voltammetry, J Am Chem Soc. 130: 6682-6683. doi: 10.1021/ja801071n
Michel LV, Ye T, Bowman, SEJ, Levin BD, Hahn MA, Russell BS, Elliott SJ, Bren KL (2007) Heme attachment motif mobility tunes cytochrome c redox potential, Biochemistry 46: 11753-11760. doi: 10.1021/bi701177j
Parks B, Vacca L, Rumberger E, Hendrickson D, Christou G (2003) Effect of mechanical stress on the line width of single photon absorptions in Mn12-acetate, Physica B 329: 1181-1182. doi: 10.1016/S0921-4526(02)02099-9
Patents:
International Application published under the patent cooperation treaty (World Intellectual Property Organization); Michel LV, Hellman J (14 May 2020) Diagnosing sepsis by detecting peptidoglycan associated lipoprotein (Pal) in urine, International Publication Number WO 2020/097499 A1.
Patent Pending: Michel LV, Hellman J (November 8, 2018) Diagnosing sepsis by detecting peptidoglycan associated lipoprotein (Pal) in urine (Application No. 62/757211).
Pichichero M, Khan MN, Kaur R, Sharma S, Casey J, Michel L (Accepted August 11, 2015) US Patent 9101568: Compositions and methods related to P6.
Selected Awards and Honors
- 2024-2025 RIT's Eisenhart Award for Outstanding Teaching
- 2024 RIT Division of Access, Engagement, and Success Faculty Beacon Award
- 2024 Presidential Awards for Excellence in Science, Mathematics and Engineering Mentoring (PAESMEM) Finalist
- 2023 ChemCUR Outstanding Mentor Award from the Chemistry Division and the Council on Undergraduate Research
- 2022 ASBMB Early Career Leadership Award
- Nominated for Isaac L. Jordan Faculty Pluralism Award, RIT (2015, 2018, 2021)
- Rochester Museum and Science Center STEM Education Award, presented to WISe (2018)
- Travel Award, 2018 Annual Biomedical Research Conference for Minority Students (2018)
- Received the COS Outstanding Contributions to the Community award, RIT (2018)
- 2017 Edwina Award, RIT, Center for Women and Gender (2017)
- Salutes to Excellence Award, from the Rochester Section American Chemical Society (2016)
- Travel Award, 2016 Annual Biomedical Research Conference for Minority Students (2016)
- COS Advancing Diversity Award, RIT (2016)
- Received INSIGHT Into Diversity’s Inspiring Women in STEM national award (2015)
- Eisenhart Award for Outstanding Teaching Finalist, RIT (2015-2016)
- College of Science Outstanding Student Mentor Award, RIT (2014)
- College of Science Leadership Award, RIT (2013)
- Featured College of Science faculty member, RIT’s Annual Faculty Scholarship Report (2013)
- College of Science Helpful Citizen Award, RIT (2012)
- Appointed to the National Institutes of Health Early Career Reviewer Program (2011)
- Invited to attend the NIH National Graduate Student Research Festival (2006)
- William F. Neuman Award, University of Rochester (2006)
- Nominated to Sigma Pi Sigma, National Physics Honor Society, Colgate University (2006)
- Elon Huntington Hooker Graduate Fellowship, University of Rochester (2005-2006)
- ACS Women Chemists Committee Travel Award, sponsored by Eli Lilly & Company (2005)
- The Leon L. Miller Graduate Fellowship, University of Rochester (2002)
- Nominated to Phi Eta Sigma, Colgate University (1999)
Grants
External:
Sartorius. Extension of previously existing agreement: Feasibility of size measurement and characterization of nanoparticles using a Sartorius virus counter (Co-PI) ($69,646) January 2025-Dec 2025.
NIH R21. Using nanopocket membranes to capture bacterial outer membrane vesicles from biofluids (MPI) ($417,032) June 2021- May 2023.
Sartorius. Extension of previously existing agreement: Feasibility of size measurement and characterization of nanoparticles using a Sartorius virus counter ($232,286) (Co-PI) Dec 2021-Dec 2022.
HHMI Undergraduate Science Education – Inclusive Excellence Grants 2017 (Co-PI). September 2017 – August 2022 ($1,000,000).
Nuclera Nucleics Ltd. An Automated Platform for Digital DNA Synthesis: Enabling low-cost production of long DNA products for synthetic biology (Co-PI) ($51,183) January 22, 2019 – January 21, 2020.
NIH R15. Phase Boundaries and Liquid Structure of Concentrated Eye Lens Protein Mixtures (Co-PI). September 2013 – August 2017 ($361,036).
The Camille and Henry Dreyfus Foundation: Special Grant Program in the Chemical Sciences. Quiet Chemistry: Working with Deaf Students in a Chemistry Research Laboratory (PI). September 2013 – December 2016 ($31,600).
Internal:
RIT: DRIG grant. Nuclear Magnetic Resonance and Light Scattering Studies of Eye Lens Protein Solutions (Co-PI). Dec 1, 2019 - Aug 31, 2021 ($13,000).
RIT: FEAD grant. Nanoscience-Enabled Antimicrobial Material (PI). July 1, 2019 – May 31, 2020 ($6,000).
RIT: PLIG. Development and Implementation of Retrieval Cues in Organic Chemistry, and Evaluation of their Effects on Knowledge Transfer (Co-PI). March 29, 2019 – August 21, 2020 ($5,000).
RIT: NIH Grant Writing Bootcamp. Implicating Pal in Gram-Negative Sepsis (PI). May 1, 2018 – August 31, 2019 ($10,000).
RIT: FEAD grant. Probing the Pal-Peptidoglycan Interactions and their Role in Pal Release (PI). September 1, 2017 – March 31, 2018 ($5,500).
RIT: ADVANCE Connect Grants Program. Formal Evaluation of WISe (PI). May 2017 – March 2018 ($5,000).
RIT: Ronald D. Dodge Memorial Faculty Grant. Funding to support a student research assistant on the Dreyfus Quiet Chemistry project (PI). 2016-2017 ($1000).
RIT: ADVANCE Connect Grants Program. Crouching Tiger, Hidden Bias (PI). June 1, 2016 – May 31, 2017 ($1,950).
RIT: ADVANCE Connect Grants Program. WISe Distinguished Speaker Series (Co-PI). June 1, 2016 – May 31, 2018 ($6,800).
RIT: FEAD grant. Implicating Dual oriented Pal in Gram-negative sepsis (PI). July 1, 2015 – January 31, 2016 ($6,000).
RIT: ADVANCE Connect Grants Program. From WISe to WISE: Networking for Women in Science and Engineering (Co-PI). June 1, 2015 – May 31, 2016 ($8,100).
RIT: ADVANCE Connect Grants Program. WISe Networking and Leadership Initiatives (PI). June 2014 – December 2014 ($10,200).
RIT: Provost’s Faculty Mentoring Grant. Series of Women Faculty Lunches/Discussions (PI). January – December 2012 ($1900).
RIT: Dean’s Research Initiation Grant. Testing for Lipoprotein Dual Orientation in the Outer Membrane of Gram-Negative Bacteria (PI). May 2012 – April 2013 ($15,000).
RIT/RGH: SEED grant. Searching for an alternative vaccine candidate for nontypable Haemophilus influenzae (PI). February 2011 – January 2012 ($20,000).
RIT: Grant Writing Boot Camp. Evaluating P6 as one of the leading vaccine candidates for Nontypable Haemophilus influenzae (PI). March 2010 – February 2011 ($5,000).
Research Students

- Nikita Robinson
- Mya Wynn
- Halley Deme
- Timur Bendlin
- Grace Jones
- Zach Hogan
- Ellen Shephard
- Abigail Pettica
- Grace Perna
- Ana Paolini Carrano
- John Fernandez
- Mallory Cooper
- Morgan Long
- Marala Perinpanayagam
- Garrett Meehan
- Kelsi Berghahn
- Sydney Maggio
- Ian McMullen
- Aaron Serrano
Biomedical Engineering Contingent: Pani Torabian

>95% of Dr. Michel’s research student graduates (not including brief interns) have or are pursuing higher degrees in science or medicine and/or are currently working in the science, research, or medical fields. Former research students are enrolled in or graduated from PhD programs, MD/DO programs, PharmD programs, Nursing programs, MS programs, PA programs, and Law School (for patent law) at some of the world’s leading universities, including Duke, Johns Hopkins, Cornell, Yale, University of Rochester, SUNY Buffalo, Case Western, Brown, UC San Diego, Penn State, University of Michigan, Mayo Clinic, The Ohio State University, Boston University, SUNY Upstate, and Albert Einstein College of Medicine. Several former research students have received NIH, NSF, or HHMI graduate research fellowships at their current institutions, and many of Dr. Michel’s research student graduates are now doing amazing work in academic research labs, government labs, and the biotech industry.
- Dr. Jennifer Milillo (2009-2011)
- Nathaniel Huddleston, MS (Summer 2010)
- Dr. Arooj Iqbal (2010-2011)
- Dr. Danielle Weekes (2010-2011)
- Dr. Breanna Kalmeta (2009-2012)
- Dr. Kyle Grimaldi (2010-2012)
- Dr. Anthony Mangan (2010-2012)
- Dr. Melody Holmquist (2011-2012)
- Dr. Joy Snyder (2009-2013)
- Rachel Schmidt (2010-2013)
- Bethany Novick, MS (2011-2014)
- Dr. John Bettinger (2011-2014)
- Dr. Victoria MacPherson (2013-2014)
- Dr. Valerie Sgheiza (2012-2015)
- Dr. Juliana Shaw (2012-2015)
- Dr. Emily Newman (2012-2015)
- Dr. David Barnard (2012-2015)
- Jeff Shaul, MS (2013-2015)
- Dr. Casey Reulbach (2013-2015)
- Dr. Sanjana Kumar (Summer 2014)
- Alexis Russell, MPH (Summer 2014)
- Rushka Kallicharan (Summer 2015)
- Kasey Morrow, RN (2013-2016)
- Angel Payan, MS (2013-2016)
- Dr. Breanne Kisselstein (2014-2016)
- Dr. Kaylee Mathews (2014-2016)
- Bethany Novick, MS (2014-2016)
- Dr. Brooke D’Arcy (2014-2017)
- Dr. Katharine Umphred-Wilson (2014-2017)
- John Zanet (2014-2017)
- Shivani Phadke, MS (2014-2017)
- Nicole Fernandez, MS (2016-2017)
- Aaron Fadden, MS/JD (2016-2017)
- Kara Farquharson, MS (2014-2018)
- Sarah Stanton, RN (2014-2018)
- Carlie McNamara (2014-2018)
- Jeanetta Pierce, MS (2014-2018)
- Mark Zavorin, MS (2016-2018)
- Grace McGinnity (2017-2018, HS intern)
- Victoria Popov (2018, One semester)
- Dr. Emma Snyder (2015-2018)
- Nicole Pannullo (2017-2018)
- Meghan O’Neil, MS (2015-2019)
- Ciara LaClair, RN (2015-2019)
- Leslie Gallardo, MS (2017-2019)
- Aaron Fadden, MS/JD (2017-2019)
- Karett Cooper (Summer 2019, HS intern)
- Julia Faraone (2016-2020)
- Abby Melake (2019-2020)
- Morgan Bauer (2016-2020)
- Sean Lewis (2016-2020)
- Xinbei Liu, MS (2016-2020)
- Zachary Ward (2018-2020)
- Kara Farquharson, MS (2018-2020)
- Olivia Fraser, MS (2018-2020)
- Daihlia Beckford, RN (2019-2020)
- Susan Cheng, PA (2019-2020)
- Eva Earnest (2018-2020)
- Niaya Jackson (2018-2021)
- Maha Khokhar (2018-2021)
- Jacey Phillips (2019-2021)
- Tyler Pugeda, MS (2019-2021)
- Wilford Burke (2019-2021)
- Nidhi Baindur (Summer 2021 Intern)
- Jasmine Mathoan (Summer 2021 Intern)
- Caitlin Shanahan (Fall 2021 Intern)
- Zach Williams (2019-2022)
- Milena Dinu (2019-2022)
- Katie O'Neill-Knasick (2020-2022)
- Janai Perdue (2020-2022)
- Jonathan Dominguez, MS (2020-2022)
- Jada Brooks (Summer 2022, HS Intern)
- Grace McGinnity (2018-2022)
- Anna Kasper (2019-2023)
- Natalie Labbe (2019-2023)
- Isabelle Pilo (2019-2023)
- Callum Smith (2020-2023)
- Danny Colton, MS (2021-2023)
- Jamie Crawford (2021-2023)
- Yasmeen Cartwright (2021-2023)
- Aoife Cannon (2021-2023)
- Alyssa Kingston (2022-2023)
- Ulysses Hampton (2022-2023)
- Saleh Almontaser (2021-2023)
- Adaeze Collins, MS (2022-2023)
- Duncan Burnside (2023)
- Jimmy Hasselbeck (2021-2024)
- Martina Videva (2021-2024)
- Gabriela Gonzalez (2021-2024)
- Navraj Singh (2021-2024)
- Nico Burgado (2022-2024)
- Tahaara Gazali (2023-2024)
- Jasmine Nichols (2023-2024)
- Aidan Miller (2021-2024)
- Mia Kushner (2023-2025)
- Guerline Guerrier (2023-2025)
- Christina Ciko, MS (2023-2025)
Michel Group Alumni
Graduates from the Michel Research Group go on to do AMAZING things! Learn more about their lives, post-RIT.
Featured Profiles on some of our Michel Group Alumni:
Arooj Iqbal '11, MS www.rit.edu/science/spotlights/med-school-prep-skills-and-experiences-outshine-your-peers

Dr. Kyle Grimaldi '12 www.rit.edu/science/spotlights/rit-md-part-team-saves-lives

Dr. Joy Snyder '13 www.rit.edu/science/spotlights/rit-pharmd-its-more-filling-prescriptions

Dr. Kaylee Mathews '16 www.rit.edu/science/spotlights/science-exploration-brown-university-researcher
Shivani Phadke '18, MS www.rit.edu/science/spotlights/biochemistry-formulating-vaccines-merck

Nicole Pannullo '18 https://www.rit.edu/news/nicole-pannullo-named-2018-goldwater-scholar

Niaya Jackson '21 www.rit.edu/science/spotlights/no-matter-where-you-start-rit-gets-you-there

Tyler Pugeda, MS '21 www.rit.edu/news/fulbright-research-scholar-tyler-pugeda-study-investigative-treatments-alzheimers-disease

Check out this video of the Michel Lab Alumni and Where They Are Now.

Teaching
Current Courses (Semester System)
- Biochemistry I
- Biochemistry II
- Biochemistry Laboratory Techniques
- Chemical Connections
- Biochemistry for Health Sciences
- Advanced Proteins
- General Analytical Chemistry
- General Analytical Chemistry Lab
- Biowriting (co-taught)
Old Courses (Quarter System)
- Biochemistry: Metabolism
- Biochemistry: Conformation and Dynamics
- Biochemistry: Laboratory Techniques
- Biochemistry Freshman Symposium
- Circular Dichroism lecture series
- NMR spectroscopy independent study
- Chemical Literature
Photos and Videos
Quick overview of the Michel Research Lab.

Dr. Michel's introduction video to her online lectures during the spring of 2020.

The killing of George Floyd and protests around the world against systemic injustice and inequality have prompted the RIT community to more closely examine its role in working to end racism. Dr. Michel participated in a conversation to reinforce that Black Lives Matter. University leadership has pledged that the conversation, no matter how uncomfortable, must continue.
Aidan Miller and Dr. Michel had the pleasure of hosting three amazing Senior Capstone students from Rochester Prep High School over winter break. These talented young women are going places! https://www.youtube.com/watch?v=LYeWdvS1a4k

Nicole Pannullo talks about her research in the Michel Lab. https://www.youtube.com/shorts/6lwh3_ZDrIU











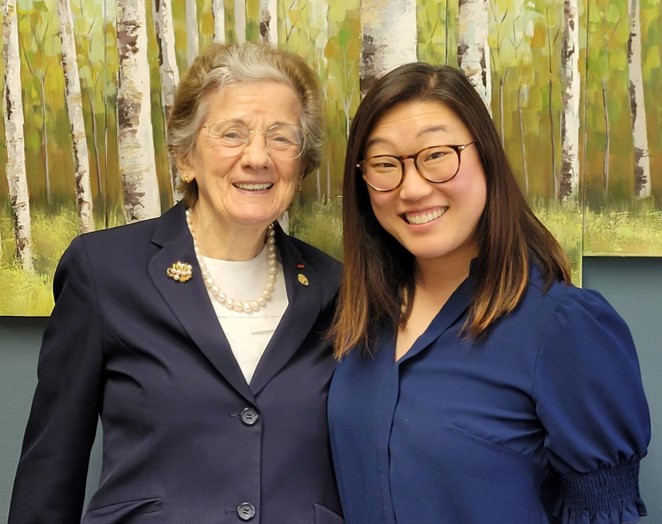
Rita Colwell and Lea Michel (May 2024)- Dr. Colwell was given an honorary degree from RIT (she has over 60 of them!).

Graduation May 2024

Michel Group 2024- End of the year Celebration!

Guerline presents her research at the NSBE Conference!
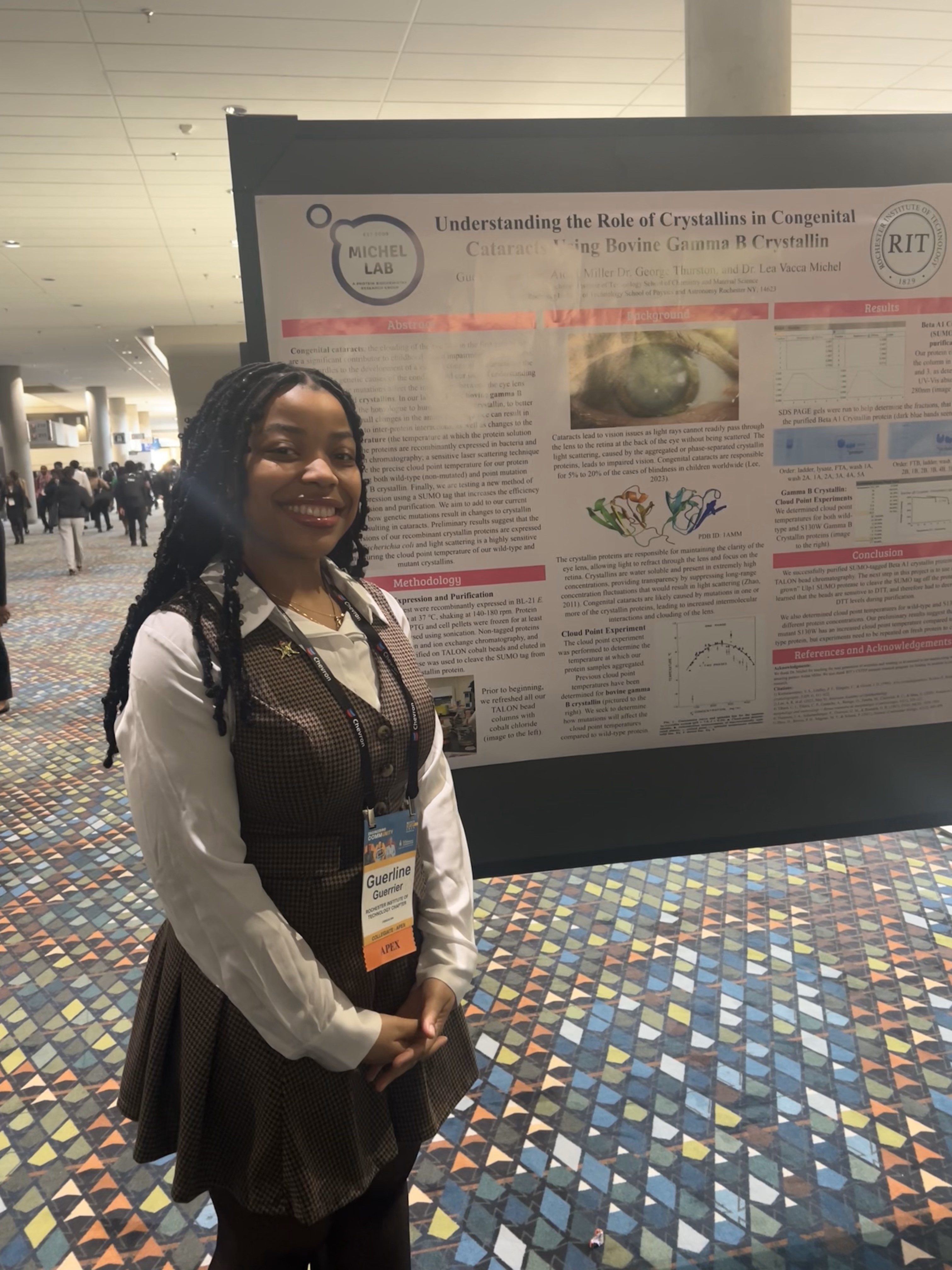

ASBMB in San Antonio March 2024!

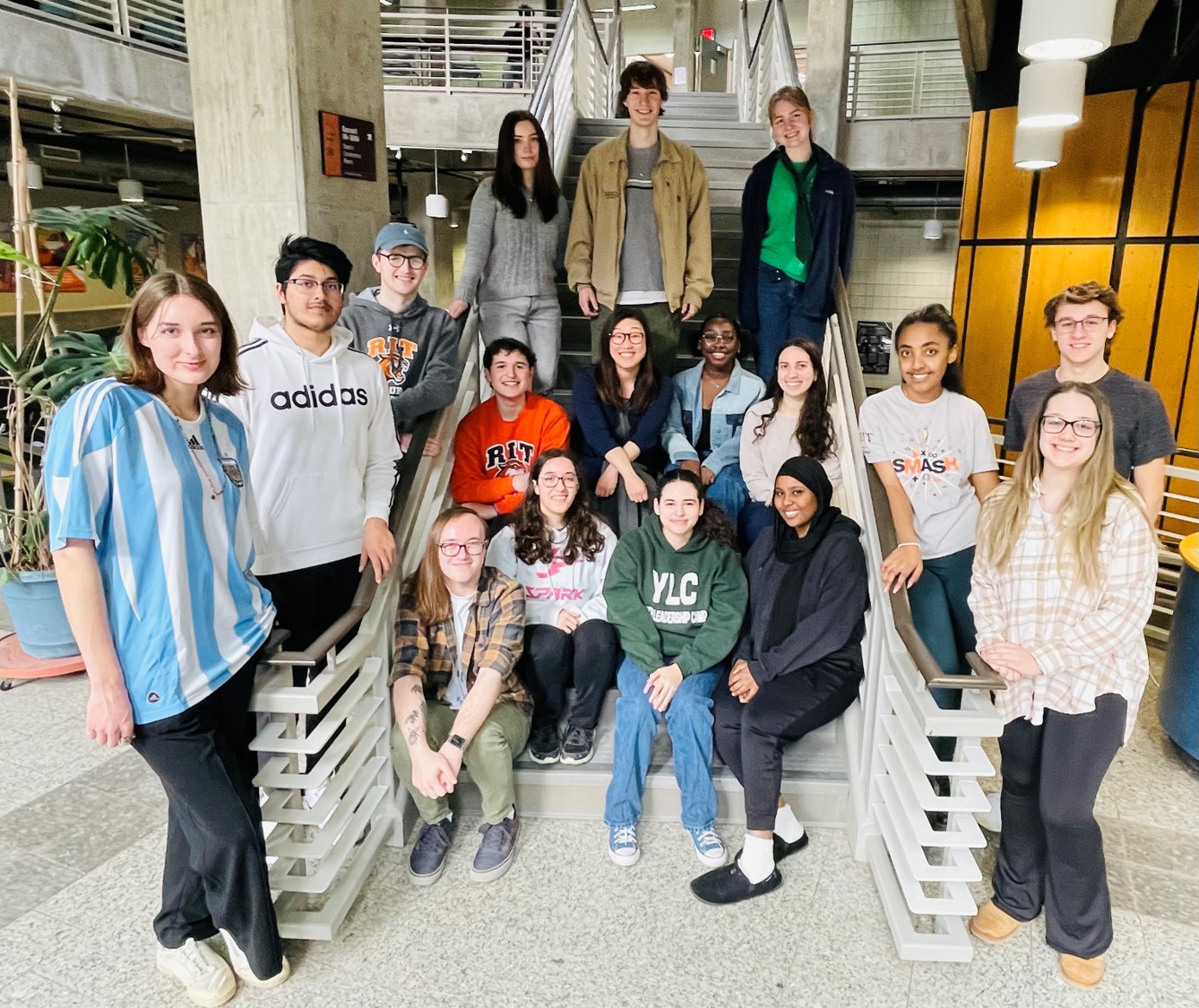
Michel group Spring 2024

Summer Research 2023 (Left to right: Alex, Navi, Martina, Lea, Guerline, Saleh, Adaeze, Nico, Aidan, Tahaara, and Gabby)!

Congratulations, Anna (the COS and RIT undergraduate commencement speaker), Isabelle, Natalie, Yasmeen, Jamie, Nico, Danny, Ulysses, Alyssa, and Aoife! So proud of our Michel research group members who graduated from RIT! From med school to PhD programs at Brown and Duke to Master's programs to scientist positions at Moderna- our graduates are off to do AMAZING THINGS!

Natalie, Isabelle, and Anna have been in the Michel lab since their first semester at RIT. We will be sad to see them leave, but they are on to exciting new adventures!
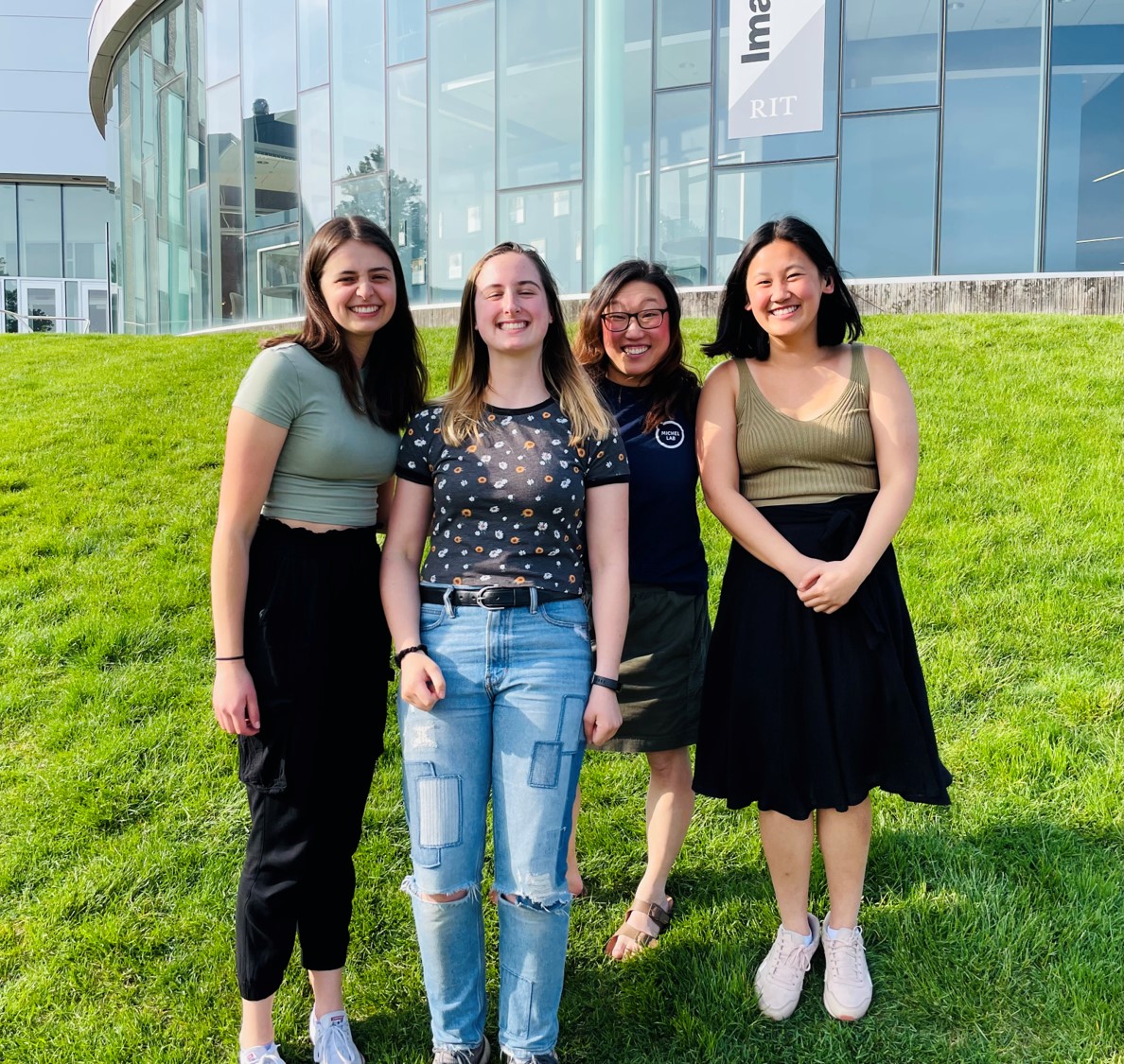

So proud of our 2023 Michel Research Student Graduates!

The Michel Lab celebrates the end of the school year with a potluck picnic!
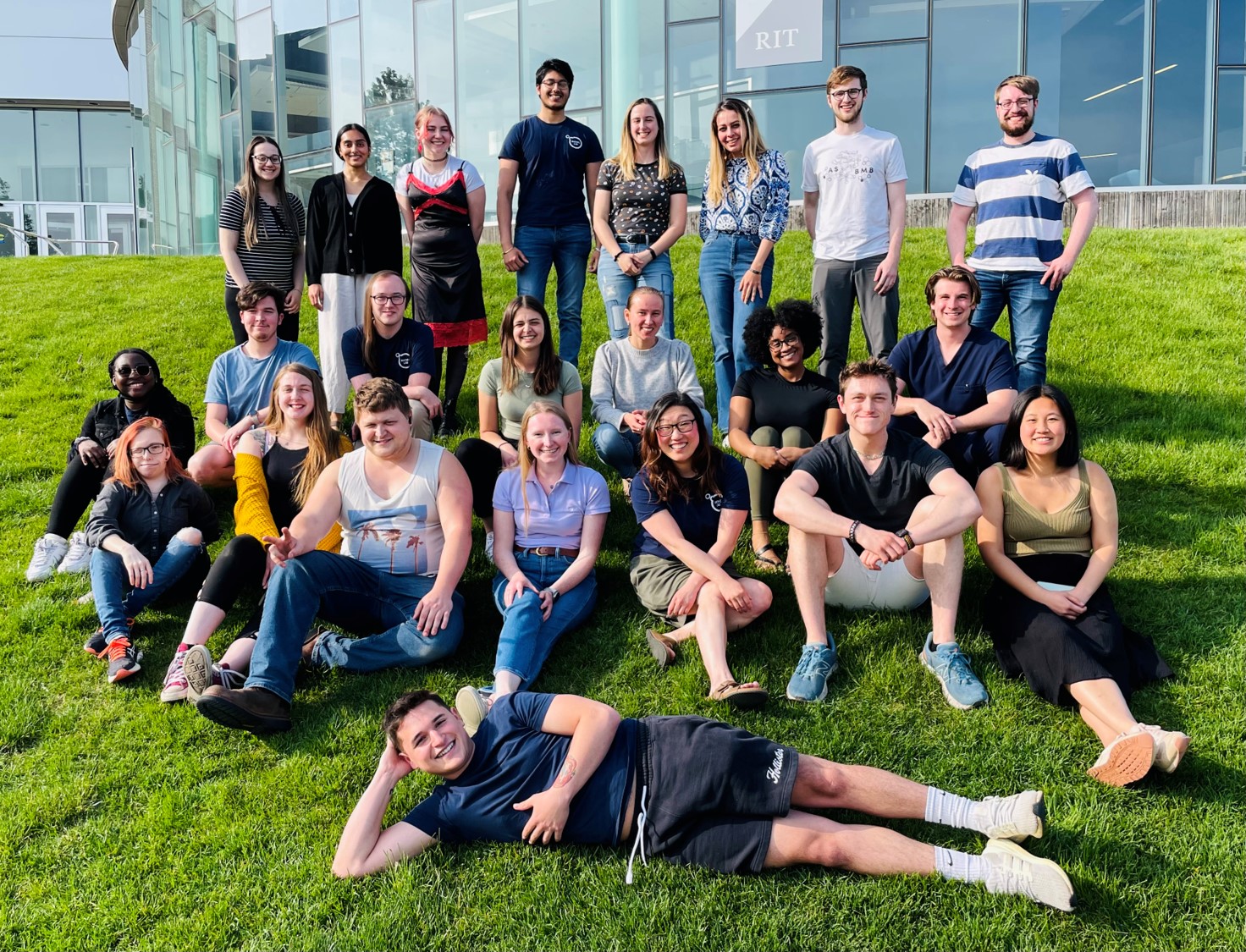

The Michel Lab represents at the national ASBMB meeting in Seattle (March 2023)
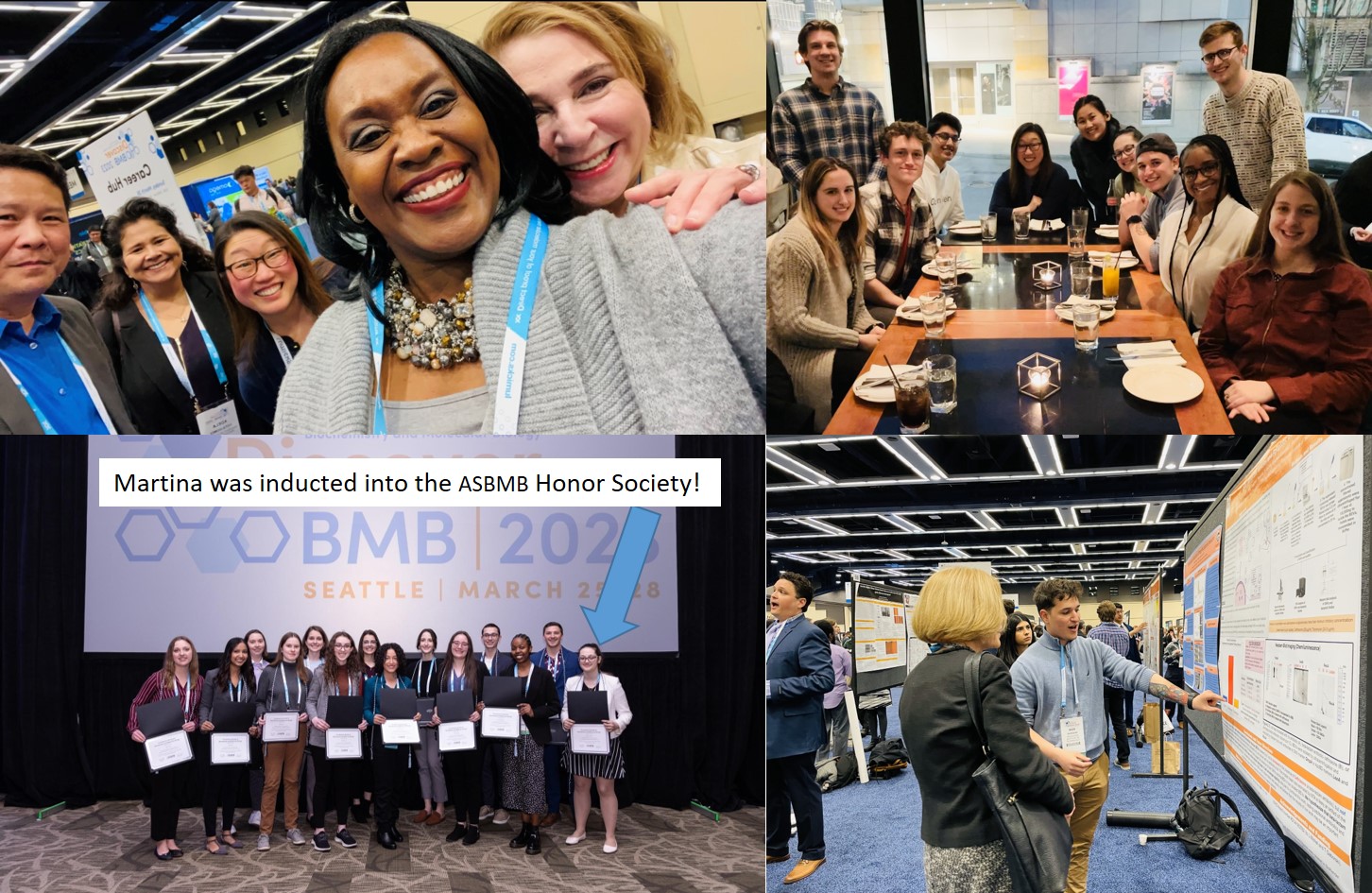 \
\
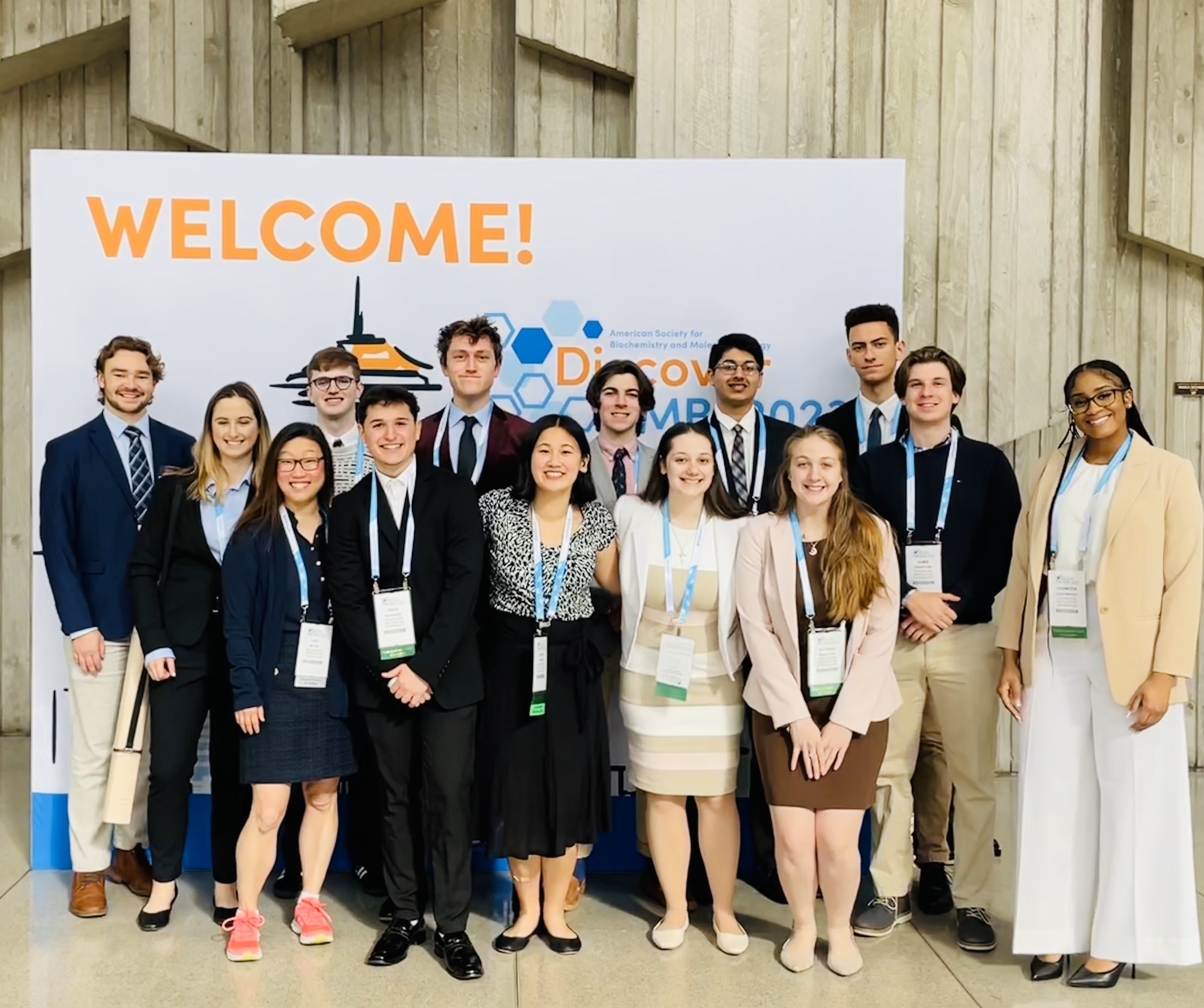

Congratulations to Anna Kasper for winning an Excellence in Student Life Award!

Congratulations to Isabelle Pilo who was selected for a research award from the Rochester Academy of Sciences!

First group meeting of 2023!

Aidan Miller and Dr. Michel had the pleasure of hosting three amazing Senior Capstone students from Rochester Prep High School over winter break. These talented young women are going places! https://www.youtube.com/watch?v=LYeWdvS1a4k
End of the Fall Semester Party (oh, and we're still missing about 1/3rd of our group)!

Summer 2022

Graduation 2022






ASBMB Meeting 2022 in Philadelphia
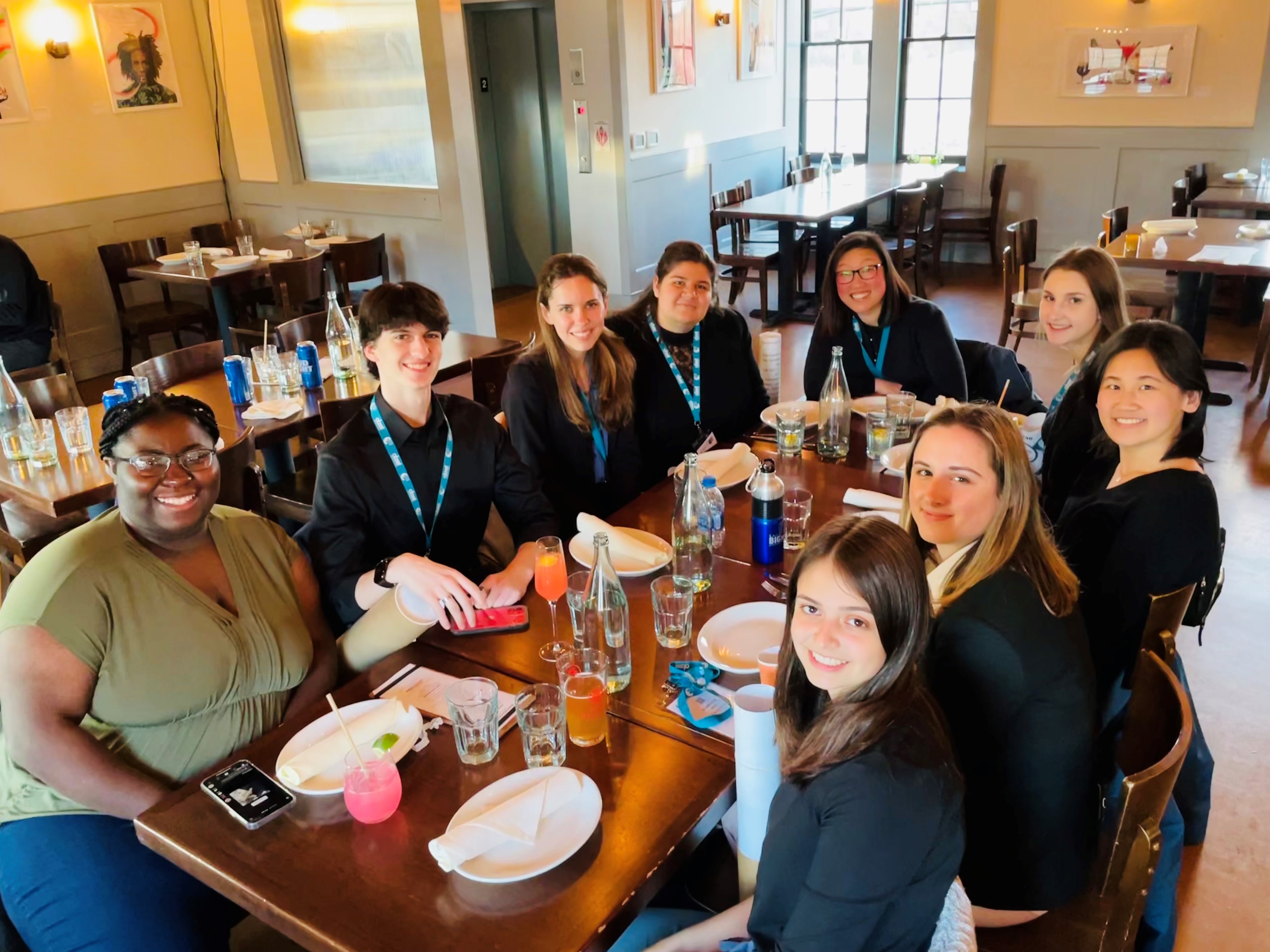
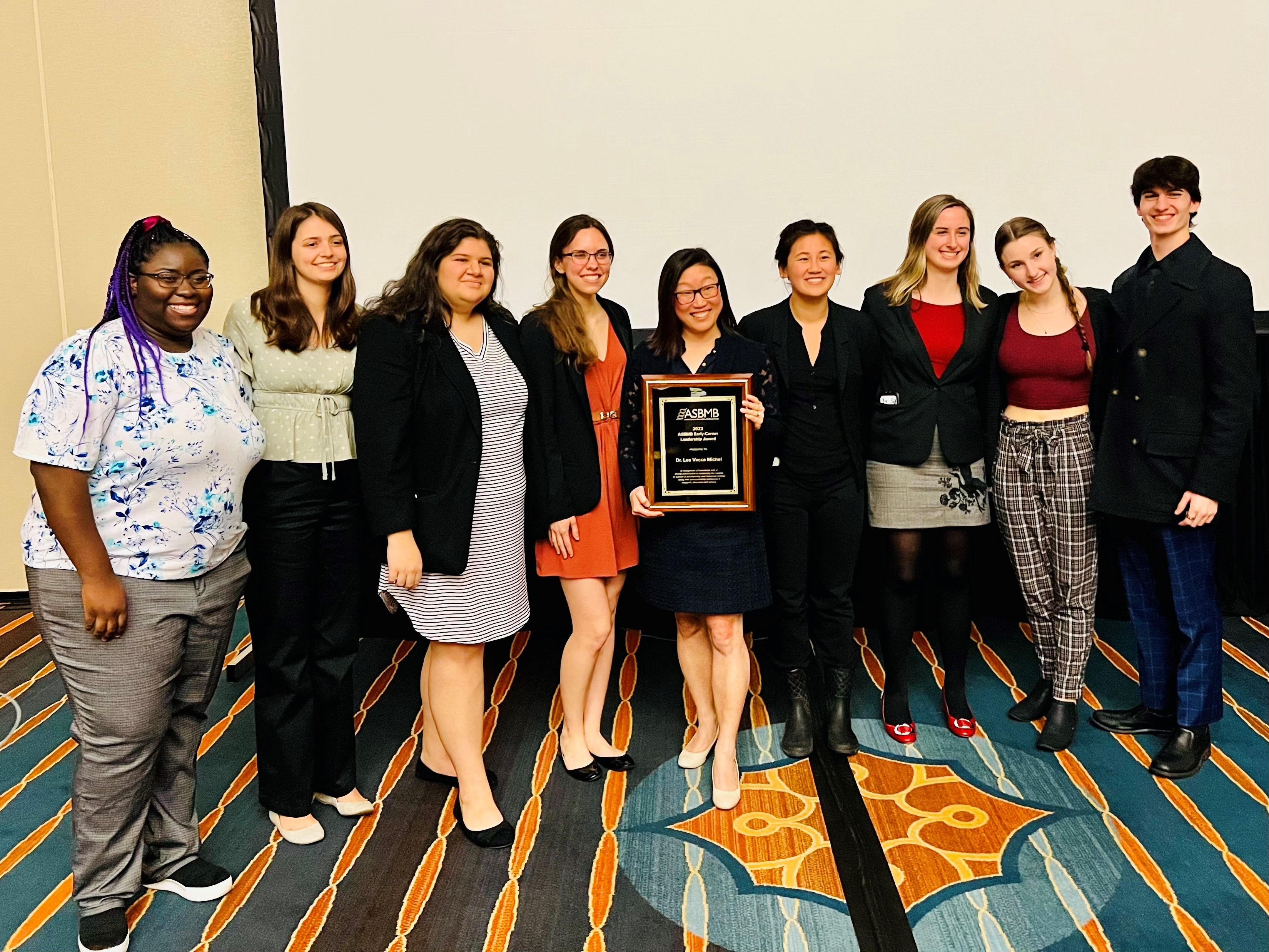
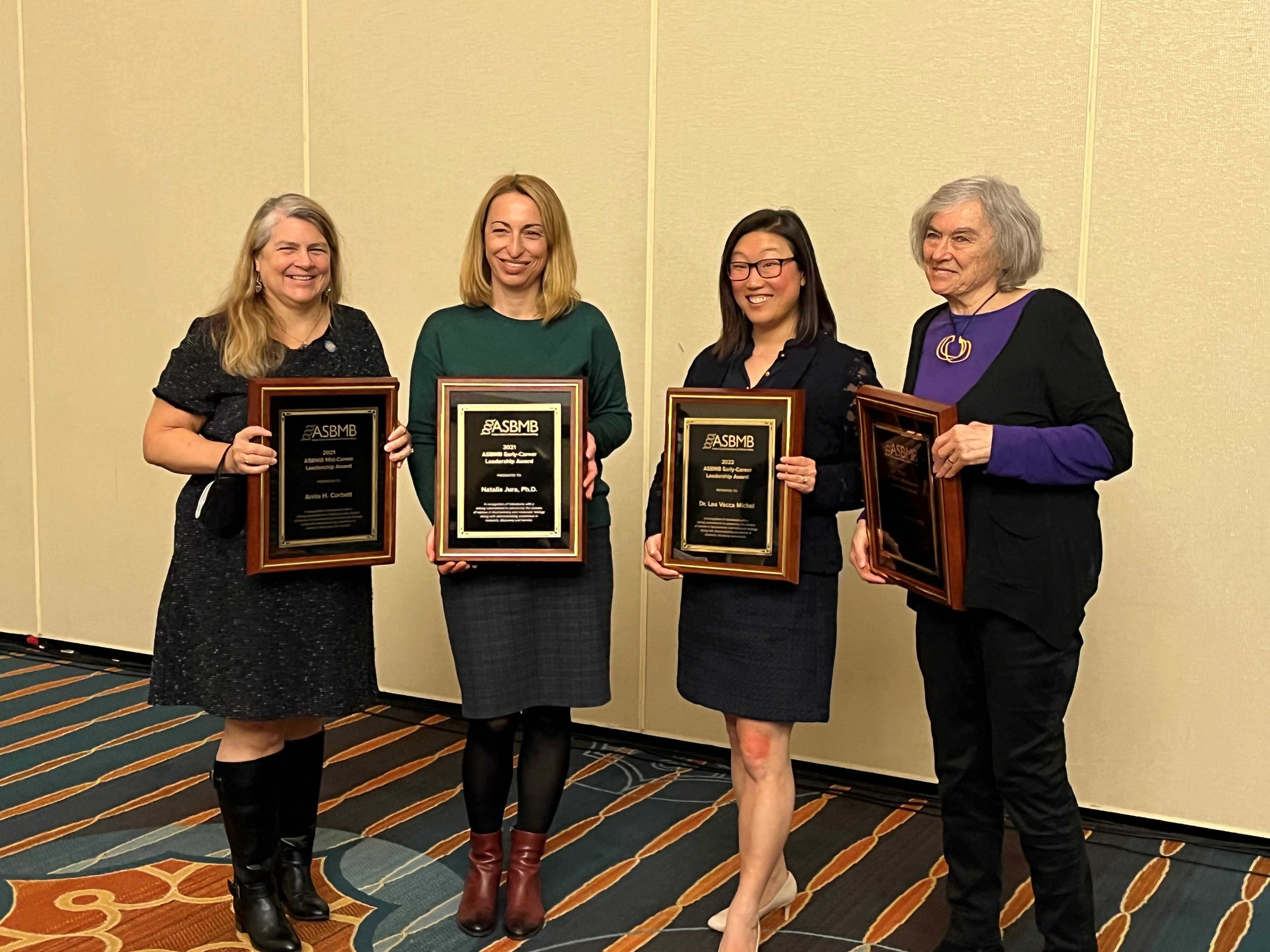
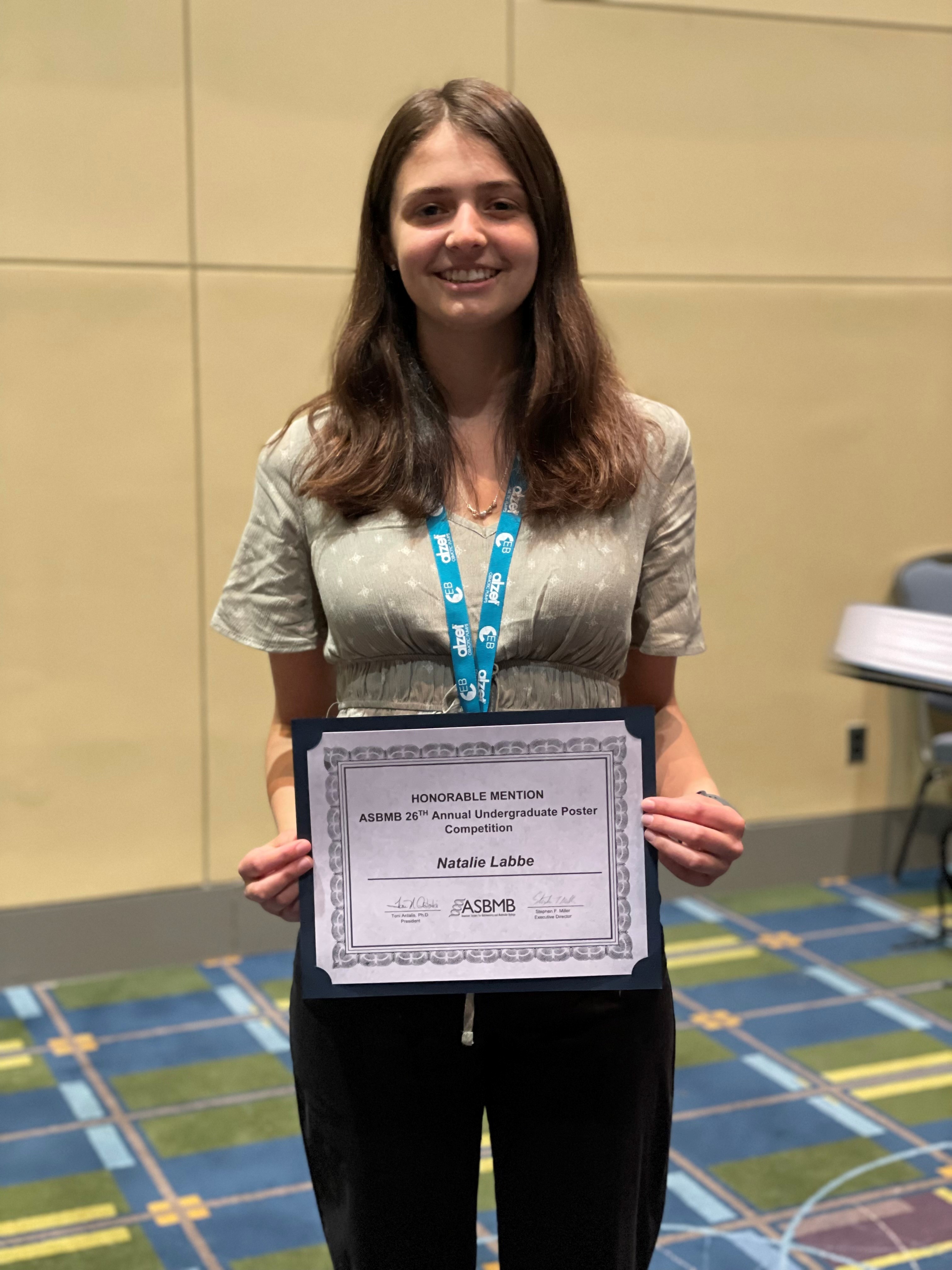
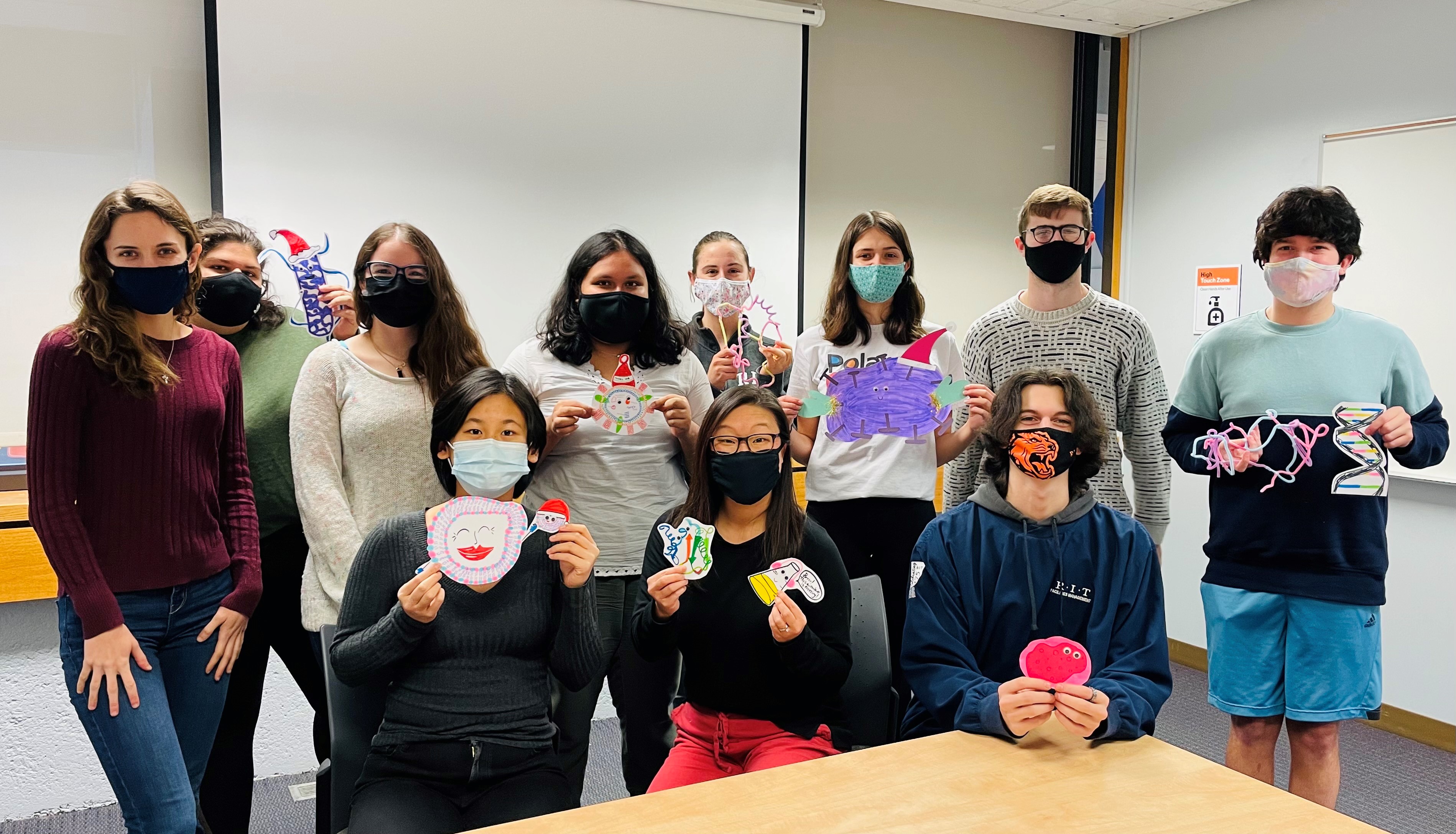








The Michel Group celebrated the end-of-the-semester with a craft session to make decorations for our lab door!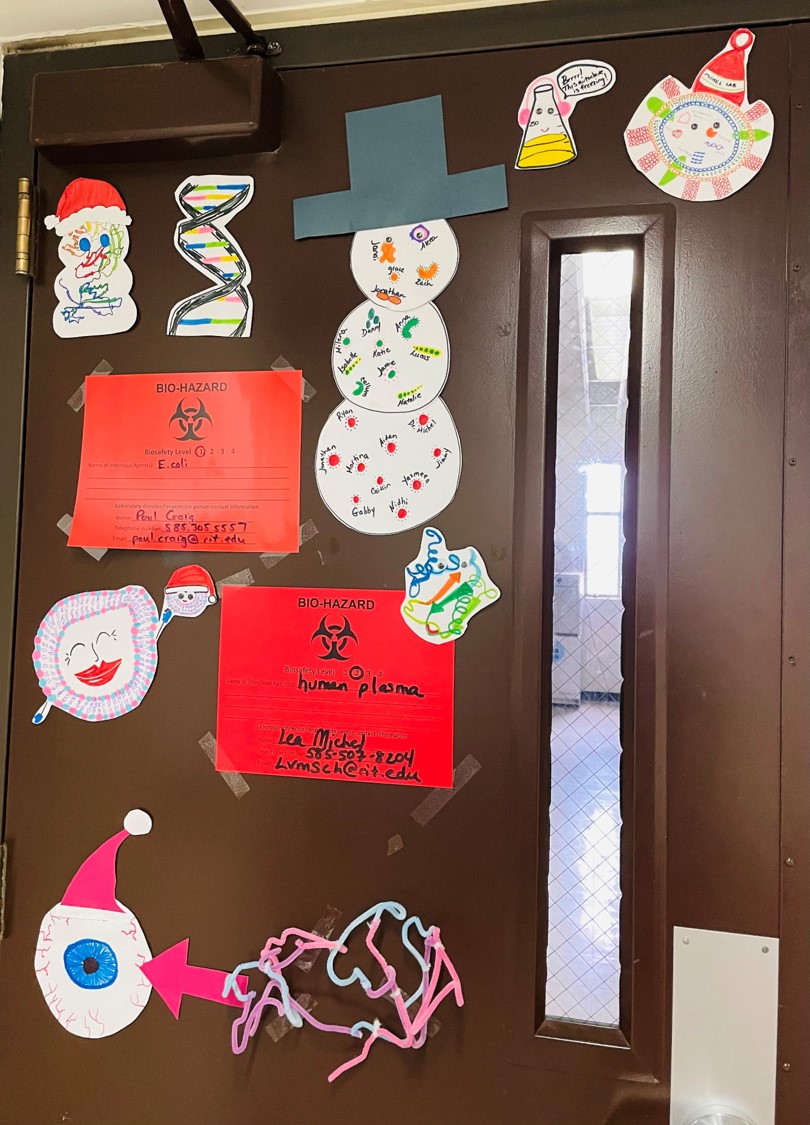
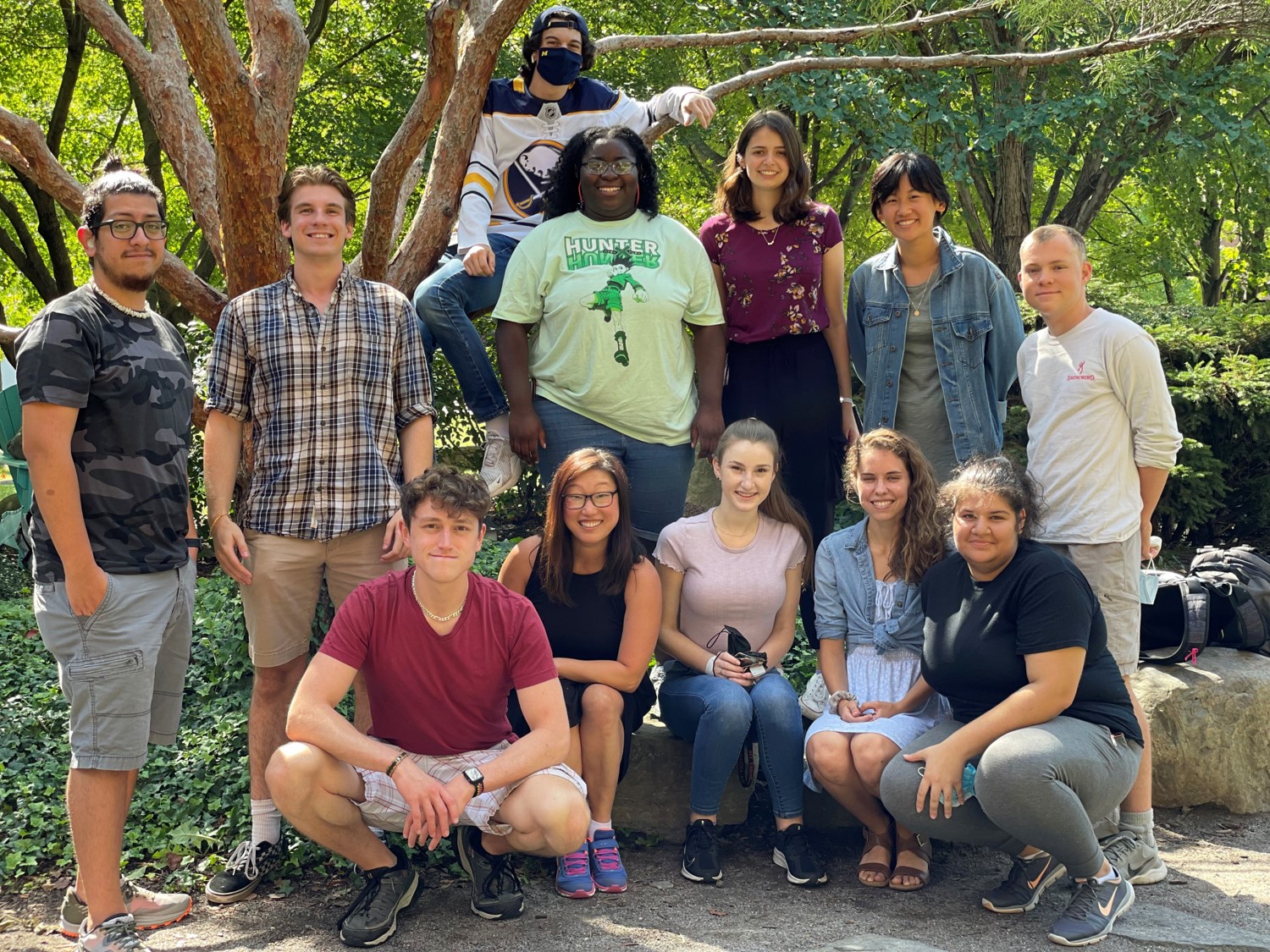


About half of the Michel Group Fall 2021
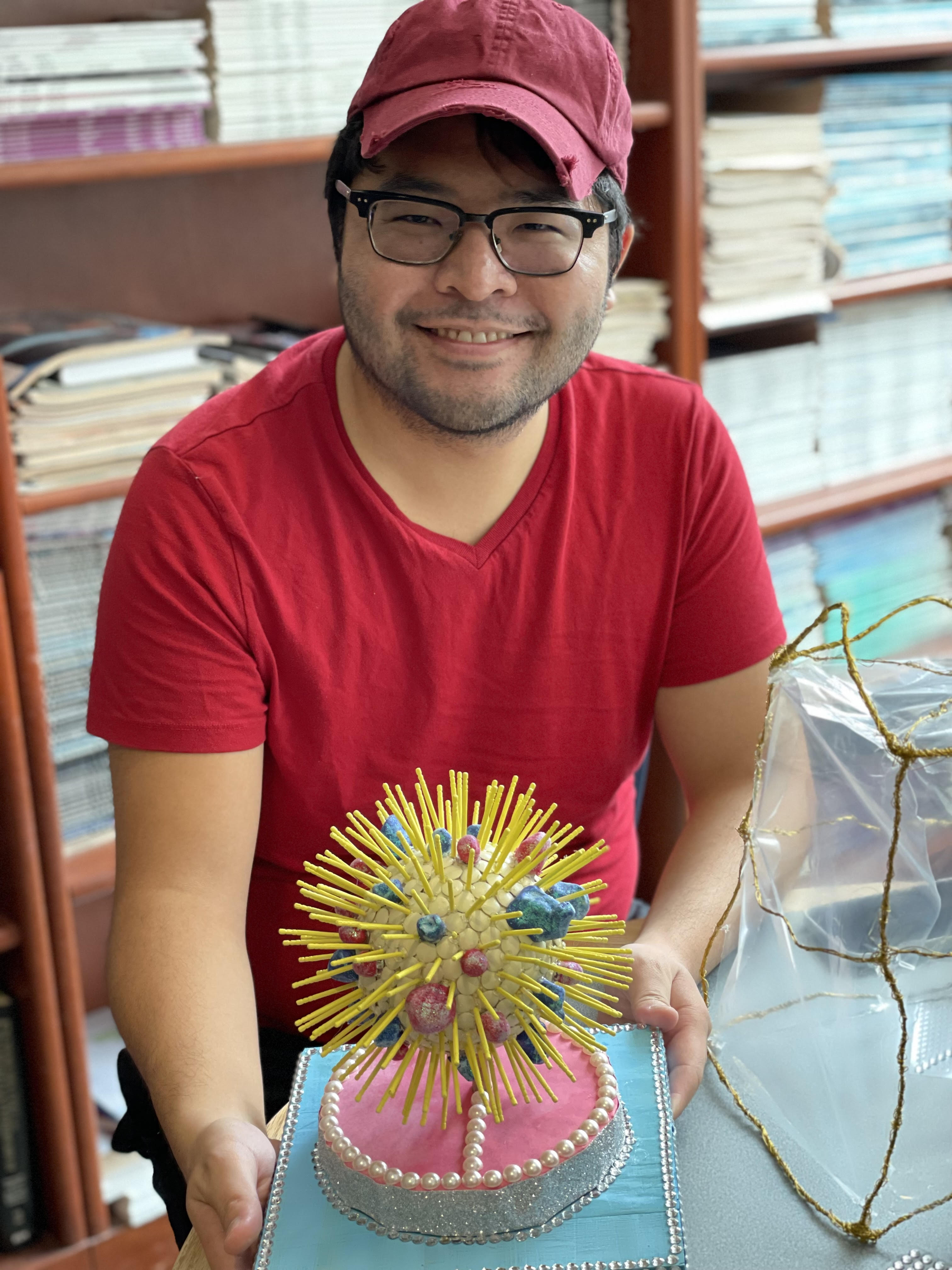

Thank you, Tyler, for your beautiful bacterial outer membrane vesicle creation!


Michel Research Group Summer 2021

Michel research group members graduate in May 2021. Congratulations, Jacey, Maha, Niaya, Wilford, and Tyler!

After more than a year, we finally got to have an in-person social event. Perfect timing to celebrate our graduating seniors (and MS grad)!

During our first (virtual) group meeting of Fall 2020, the group created this Wordle for our lab. Note, the diet (Dr) pepper (my students know me so well). Not sure about "poisoning," but I was happy to see the words "inclusive" and "welcoming."
Group meetings in the spring of 2020 were very different from group meetings of the past. Here is a photo from our last group event of the semester on Zoom.

Michel Lab Alumna, Katharine Umphred-Wilson ('17), is not only a talented scientist (PhD Program in Biomedical Sciences, Case Western University), but also a very talented artist. She made Dr. Michel's vision come true when she created this beautiful drawing for the Michel Lab 2020 Graduates.

Here is Dr. Michel's poem "Oh the Places You'll Go: RIT Chemistry Edition" that she wrote (adapted from the Dr. Seuss poem) and dedicated to the RIT School of Chemistry and Materials Science Class of 2020.
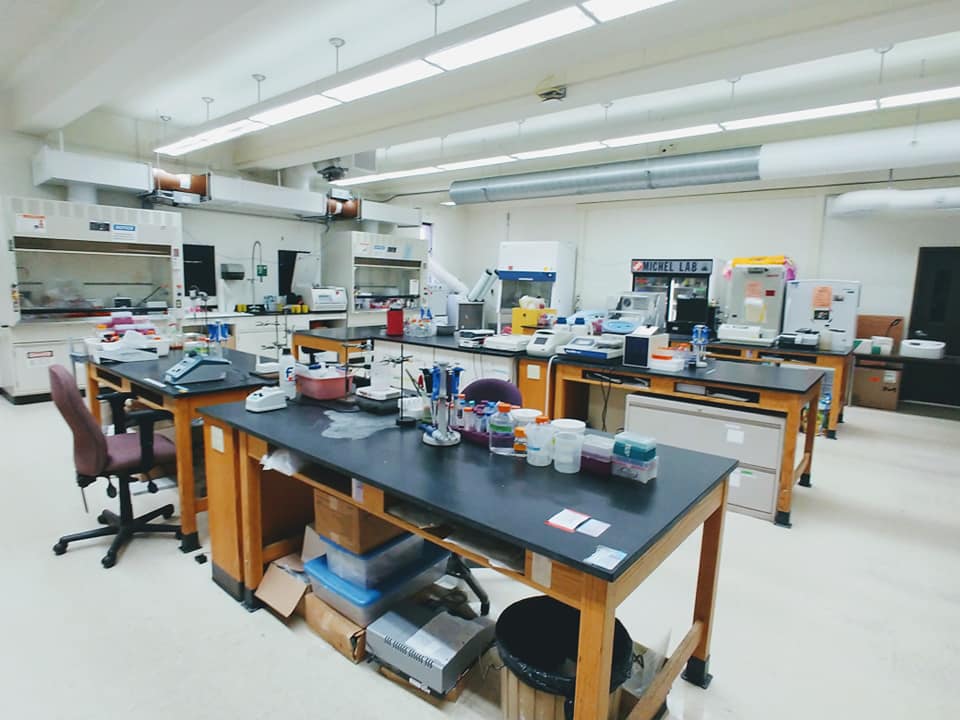

In March 2020, we closed down the Michel Lab due to the COVID-19 pandemic. It was a sad and very quiet day in the lab.
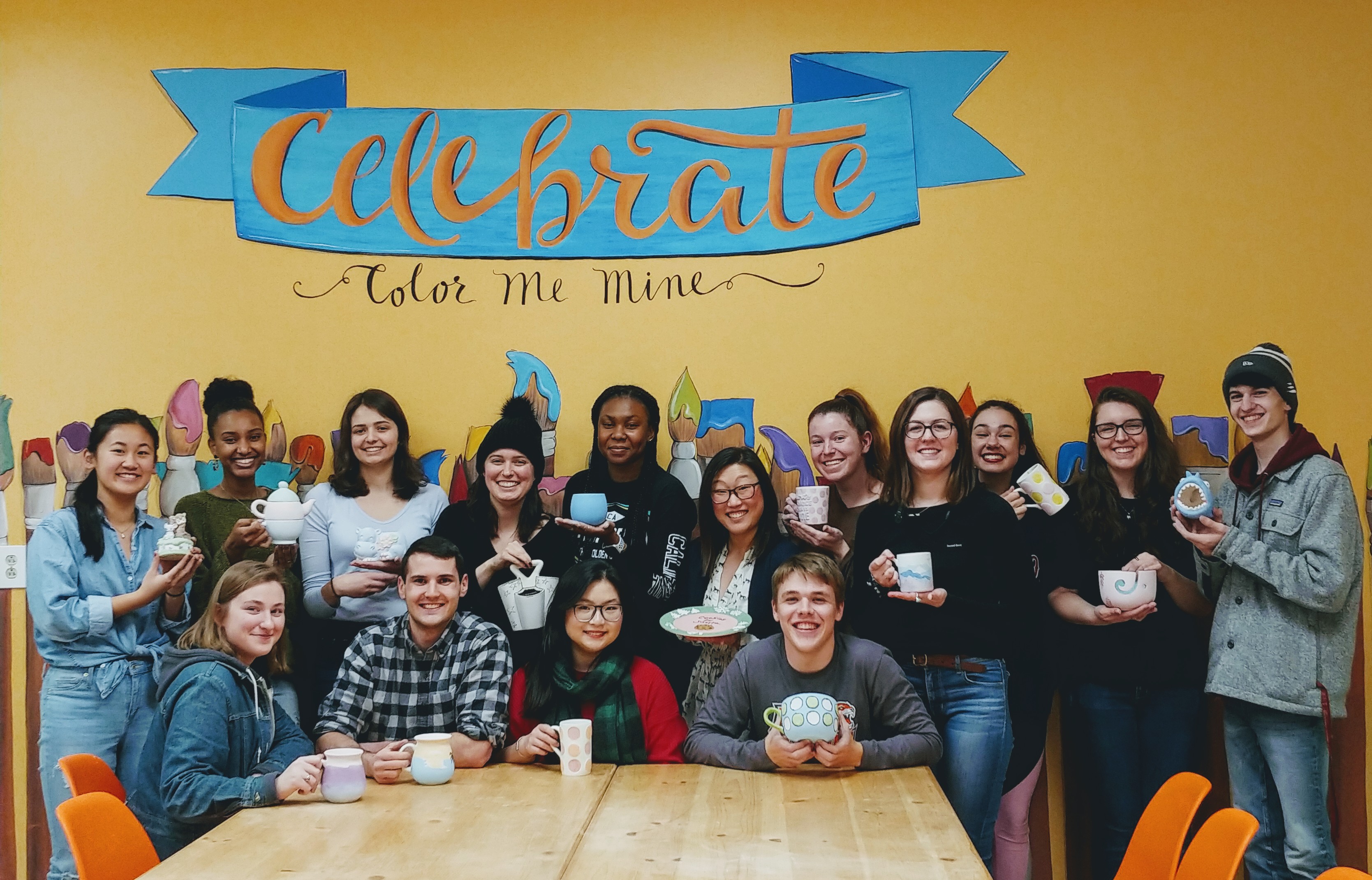

The Michel Research Group celebrates the end of the fall semester with a holiday party at Color Me Mine (we found out that several Michel group members are excellent artists)!
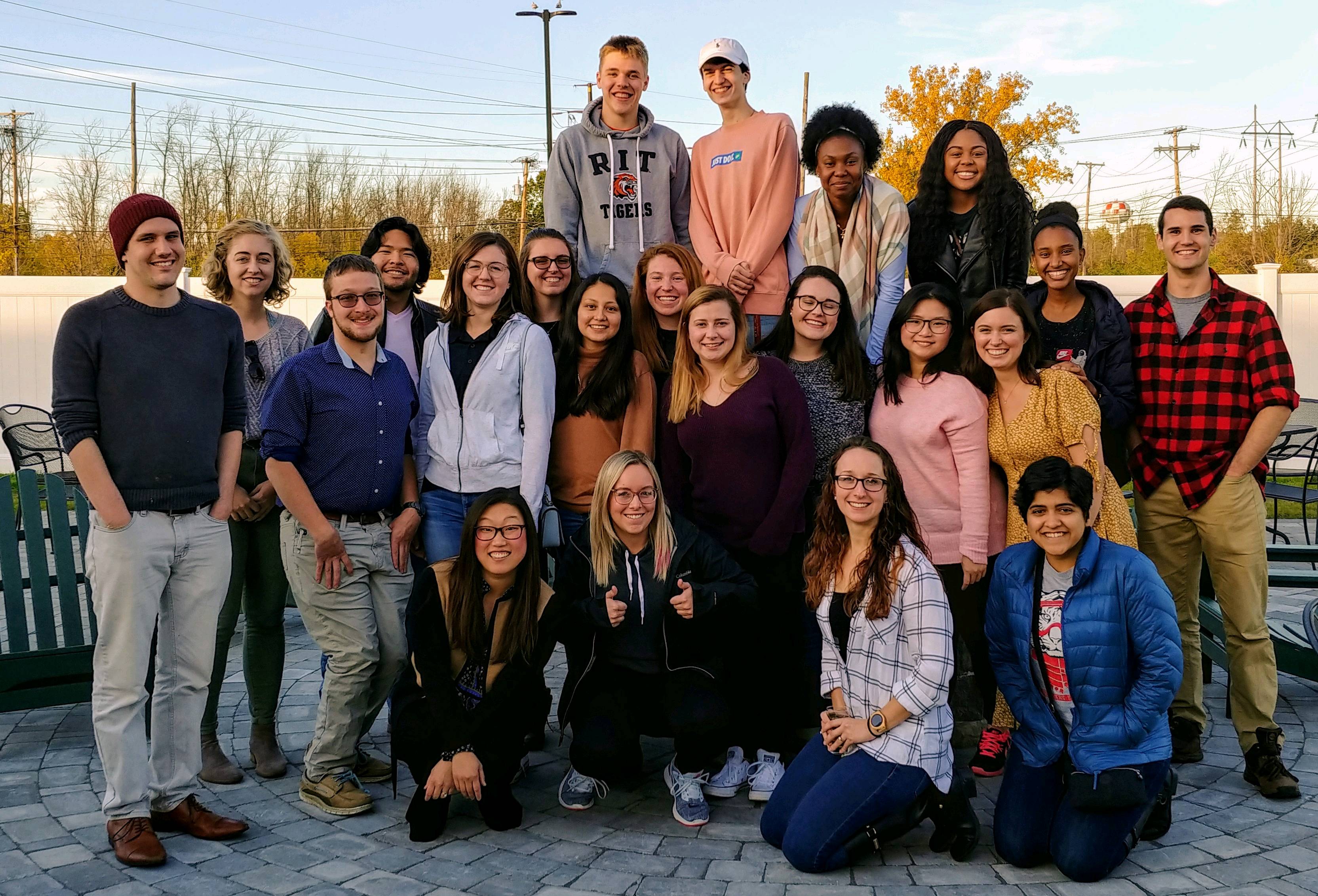

Michel Research Group 10 Year Reunion Party! Current and former research group members celebrate 10 years of the Michel lab!
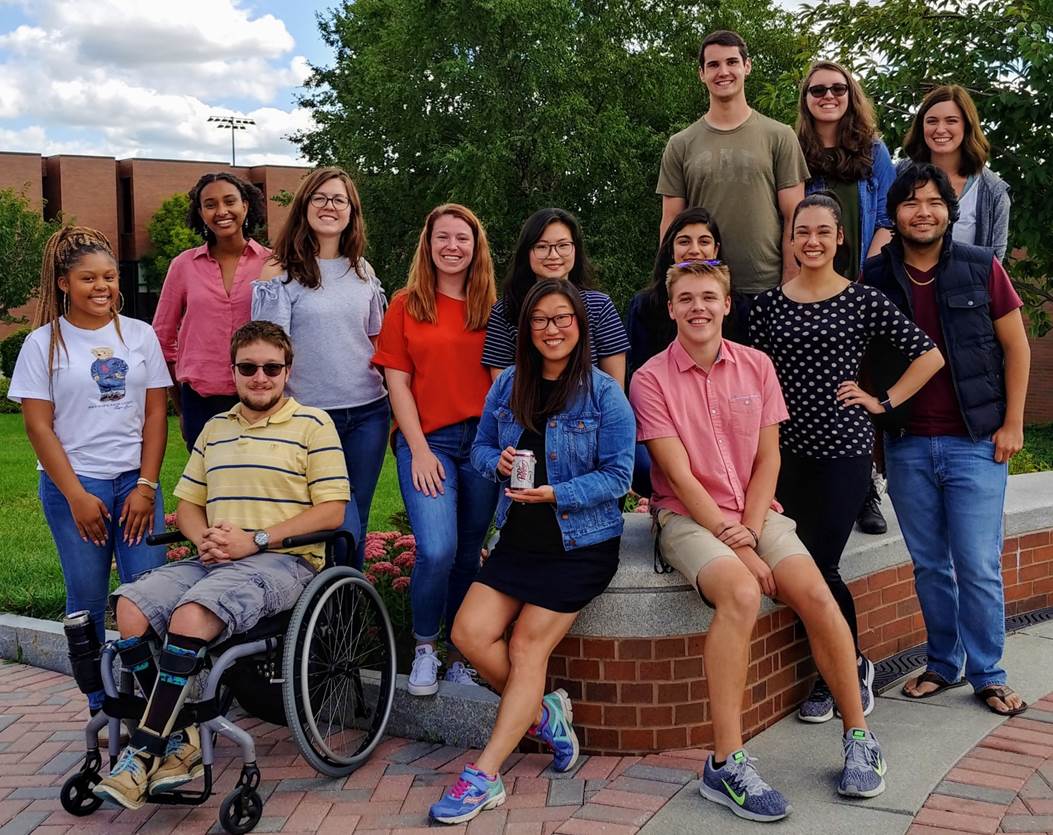

Group photo (October 2019)
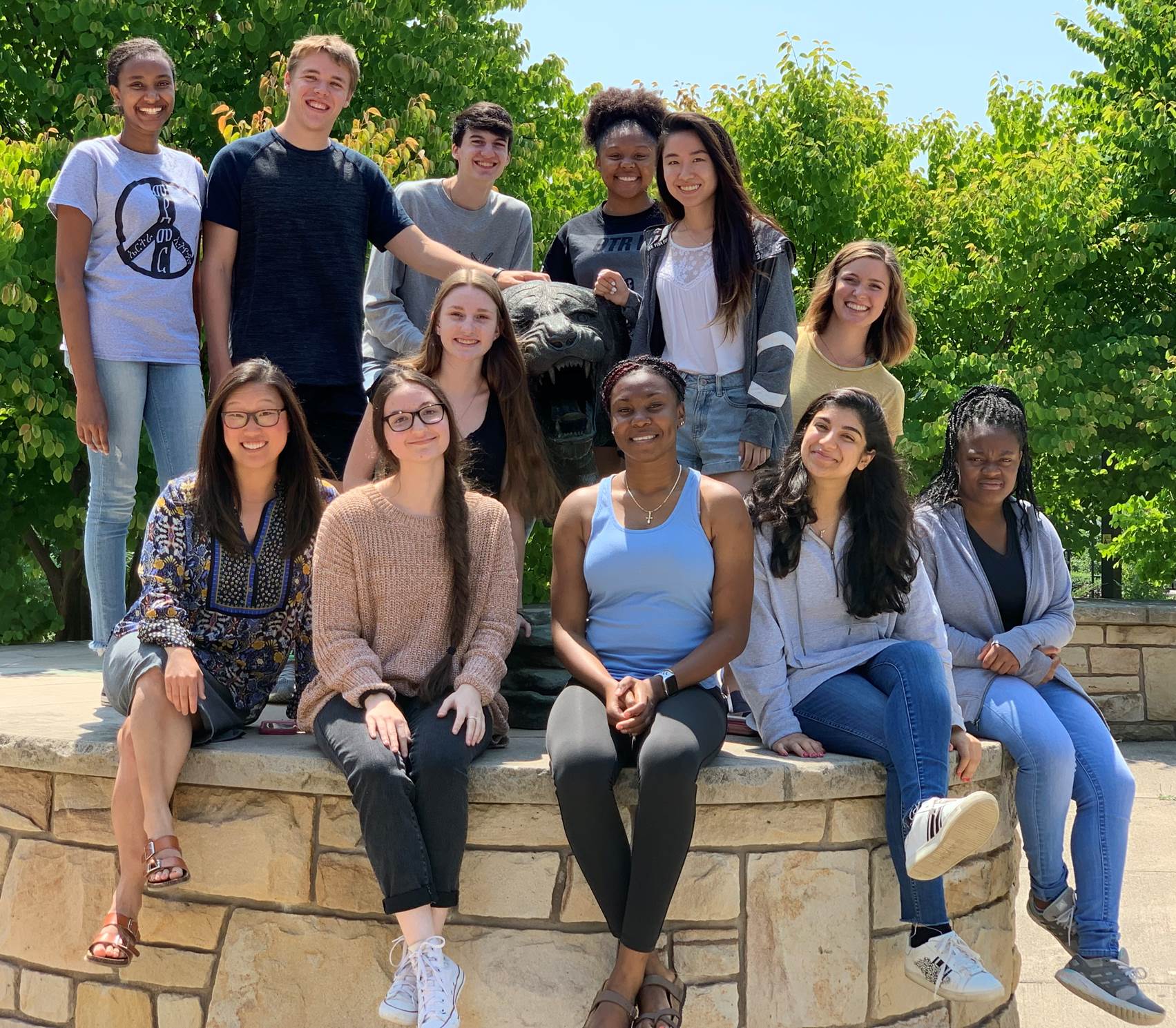

Michel Research Group- Summer 2019
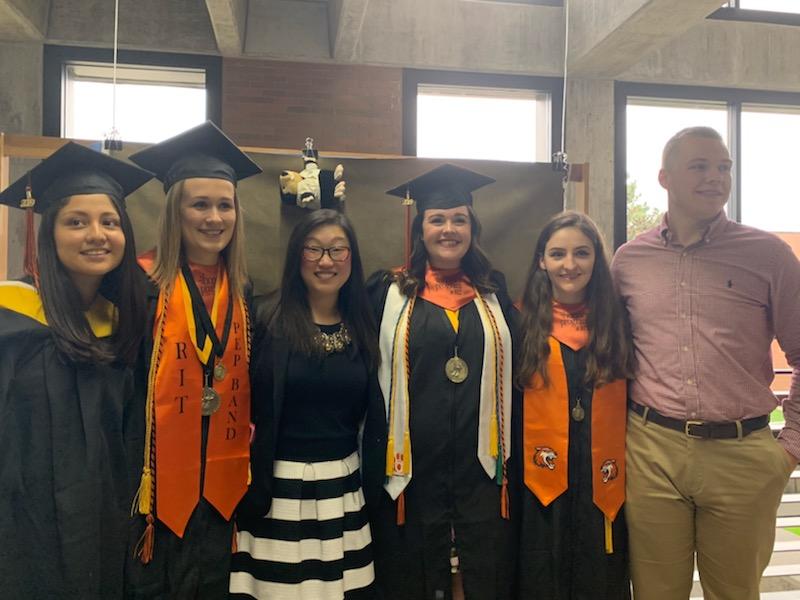

Michel Research Group Graduates 2019
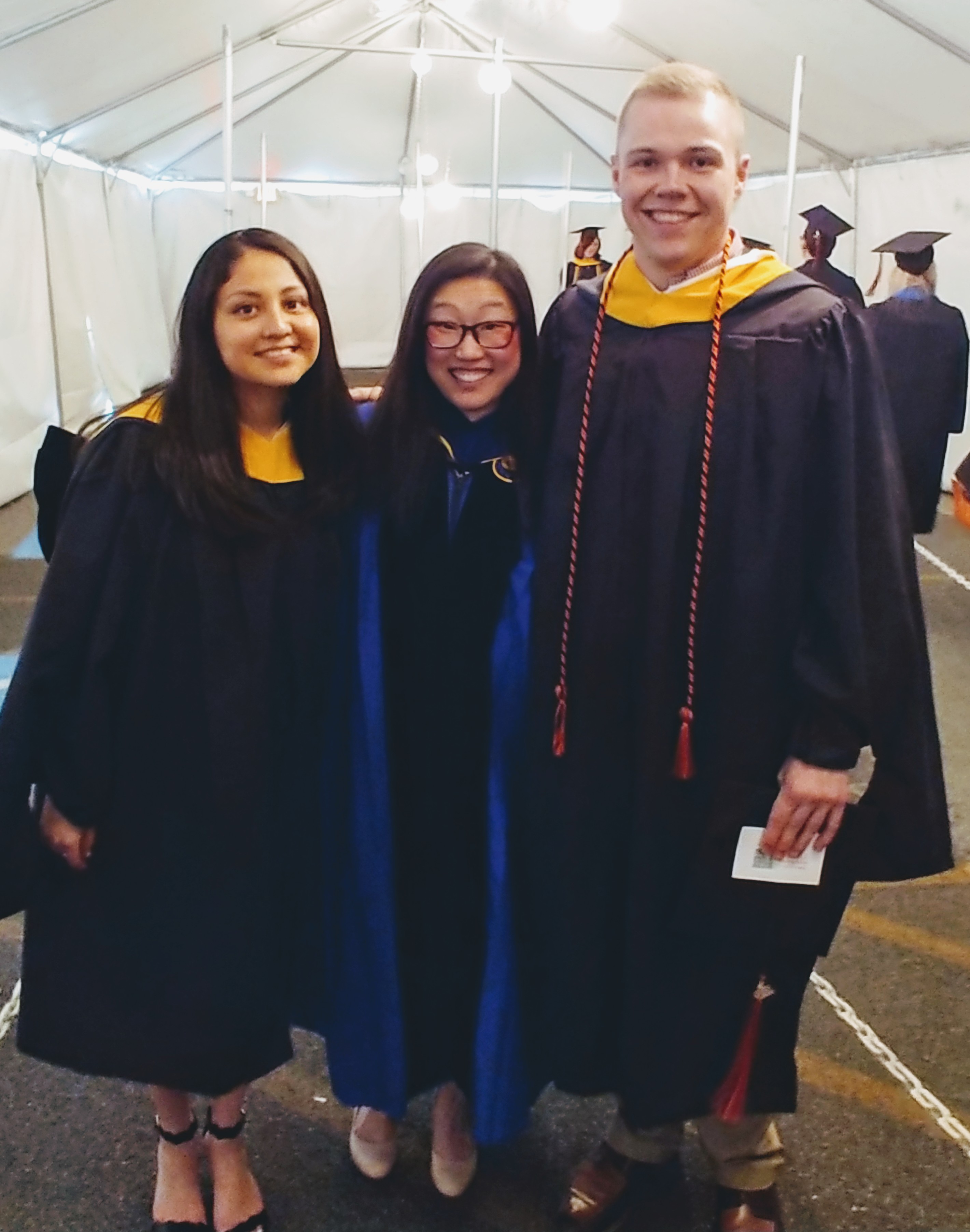

Masters Graduates 2019
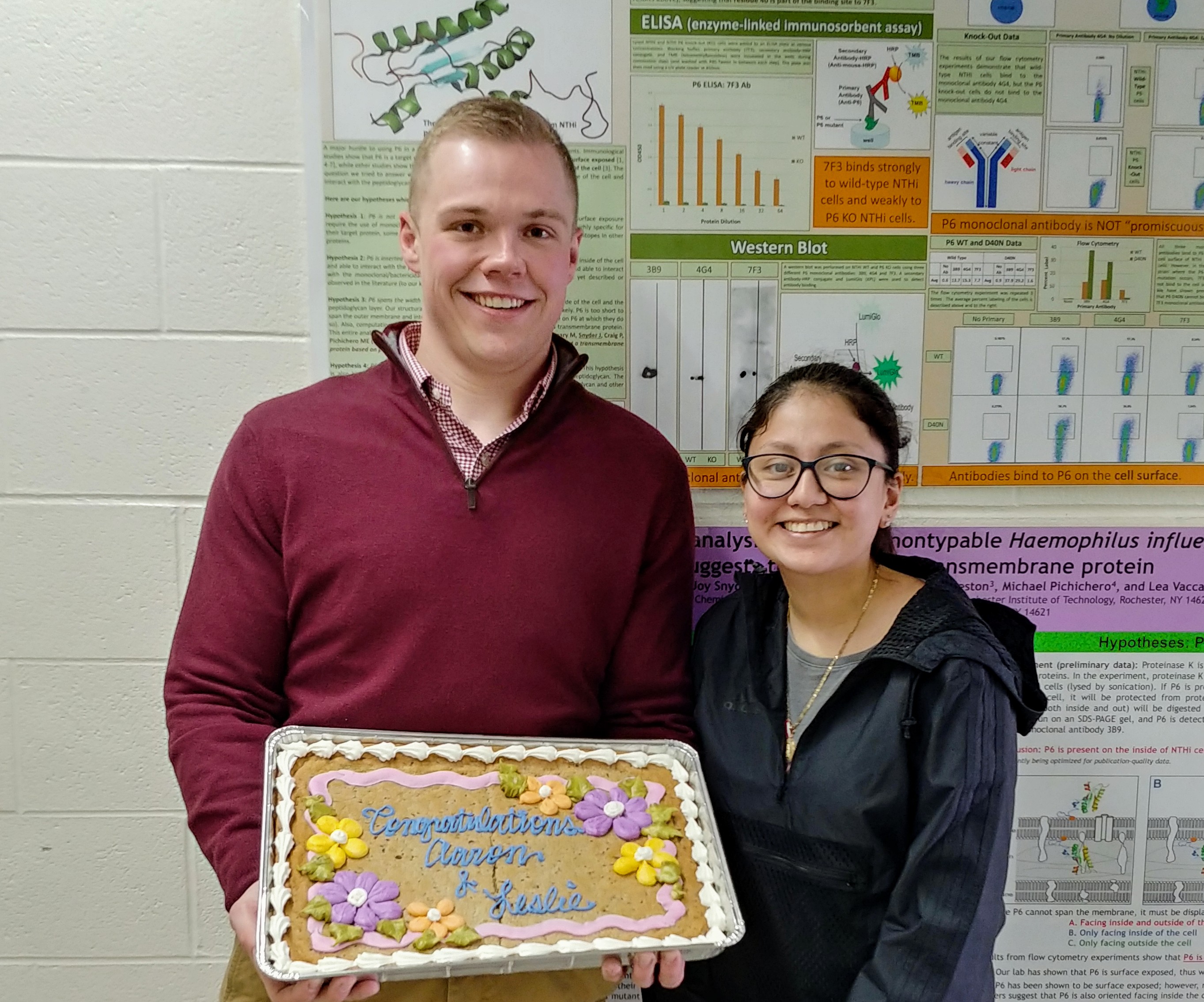

Leslie and Aaron give their MS thesis presentations
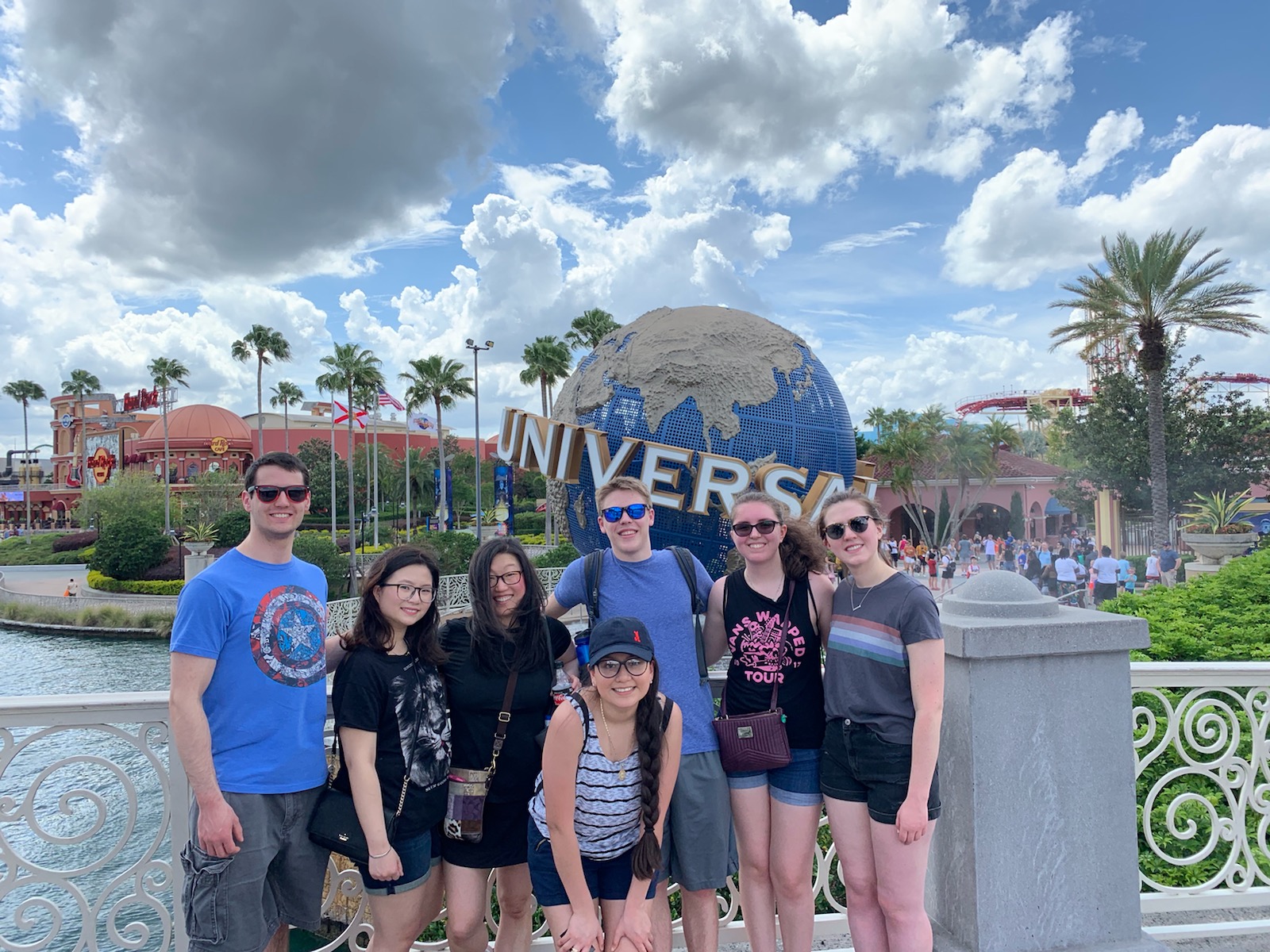

Michel Group at Universal Studios (ASBMB meeting 2019)
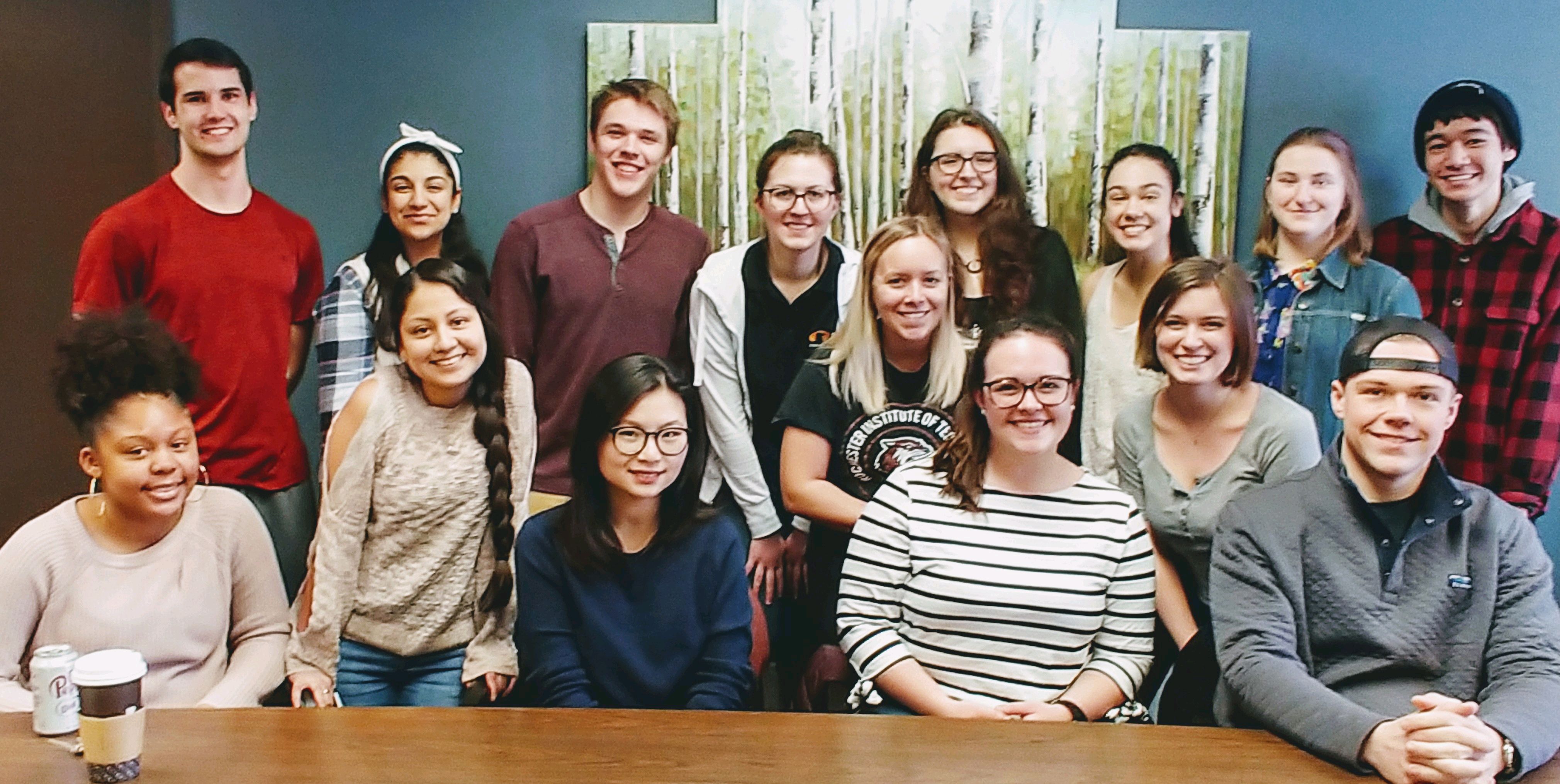

Michel Research Group Photo (March 2019)
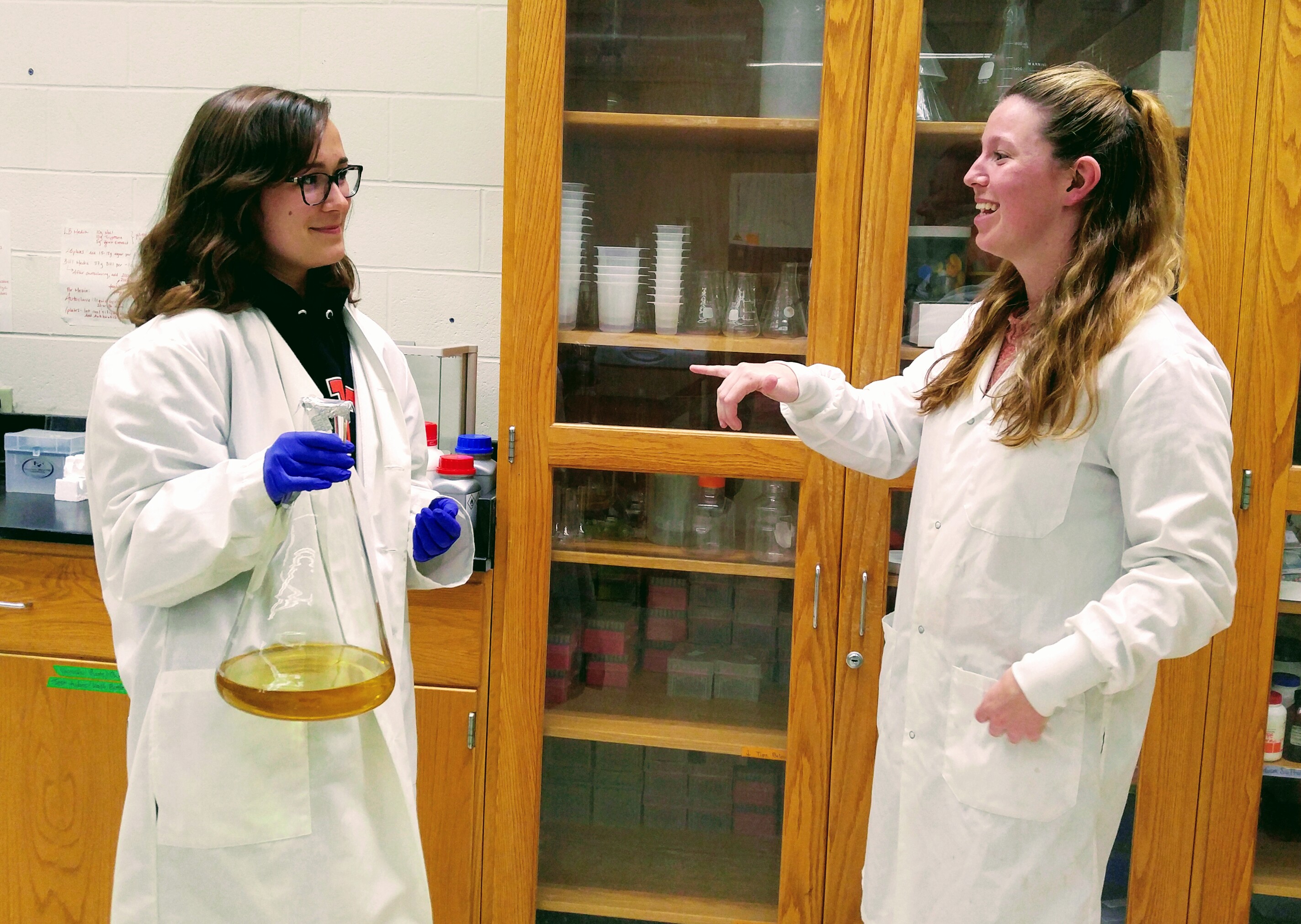

Nicole and Kara signing in the lab
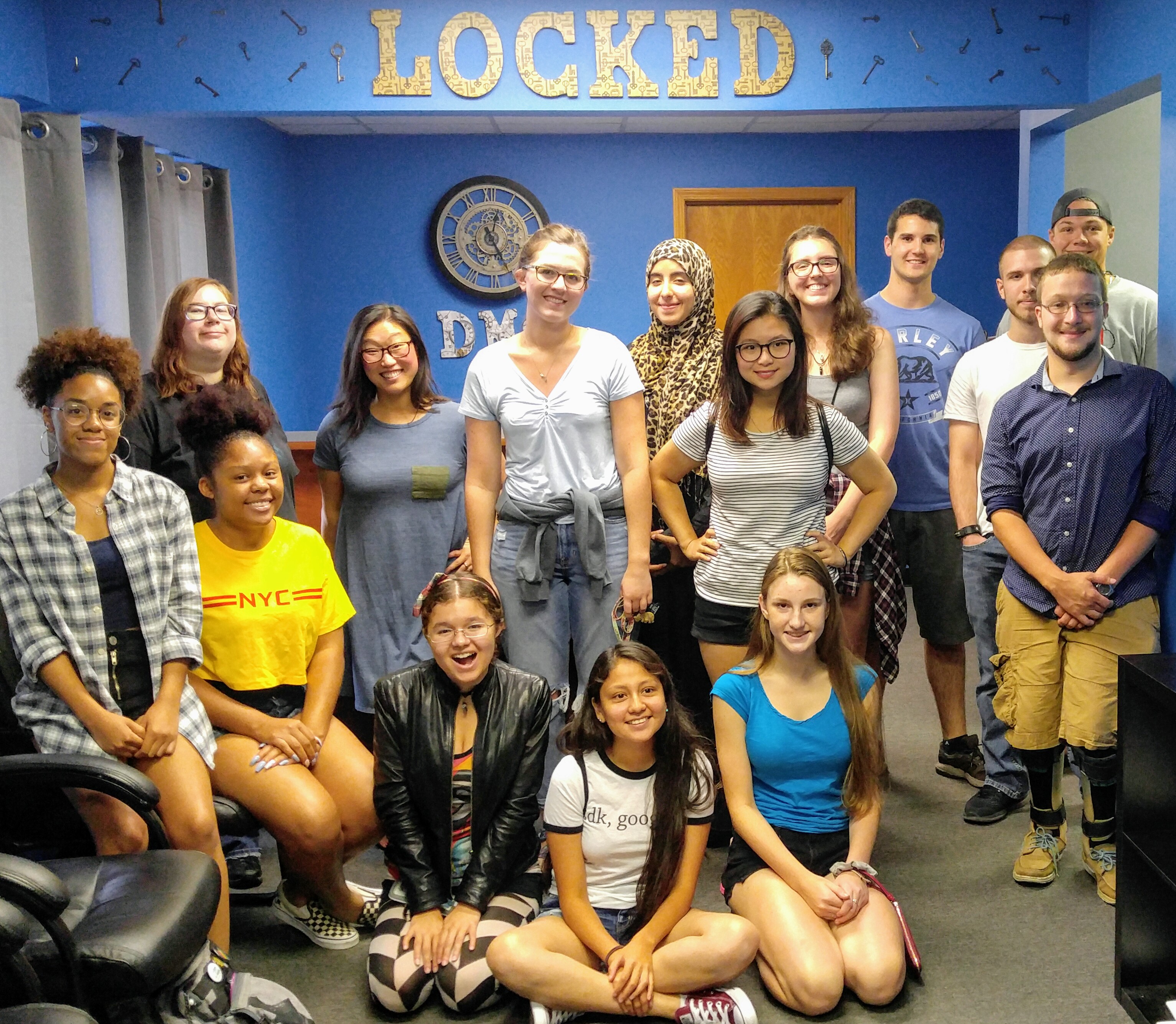

Michel Research Group and Inclusive Excellence students can't escape the Locked room
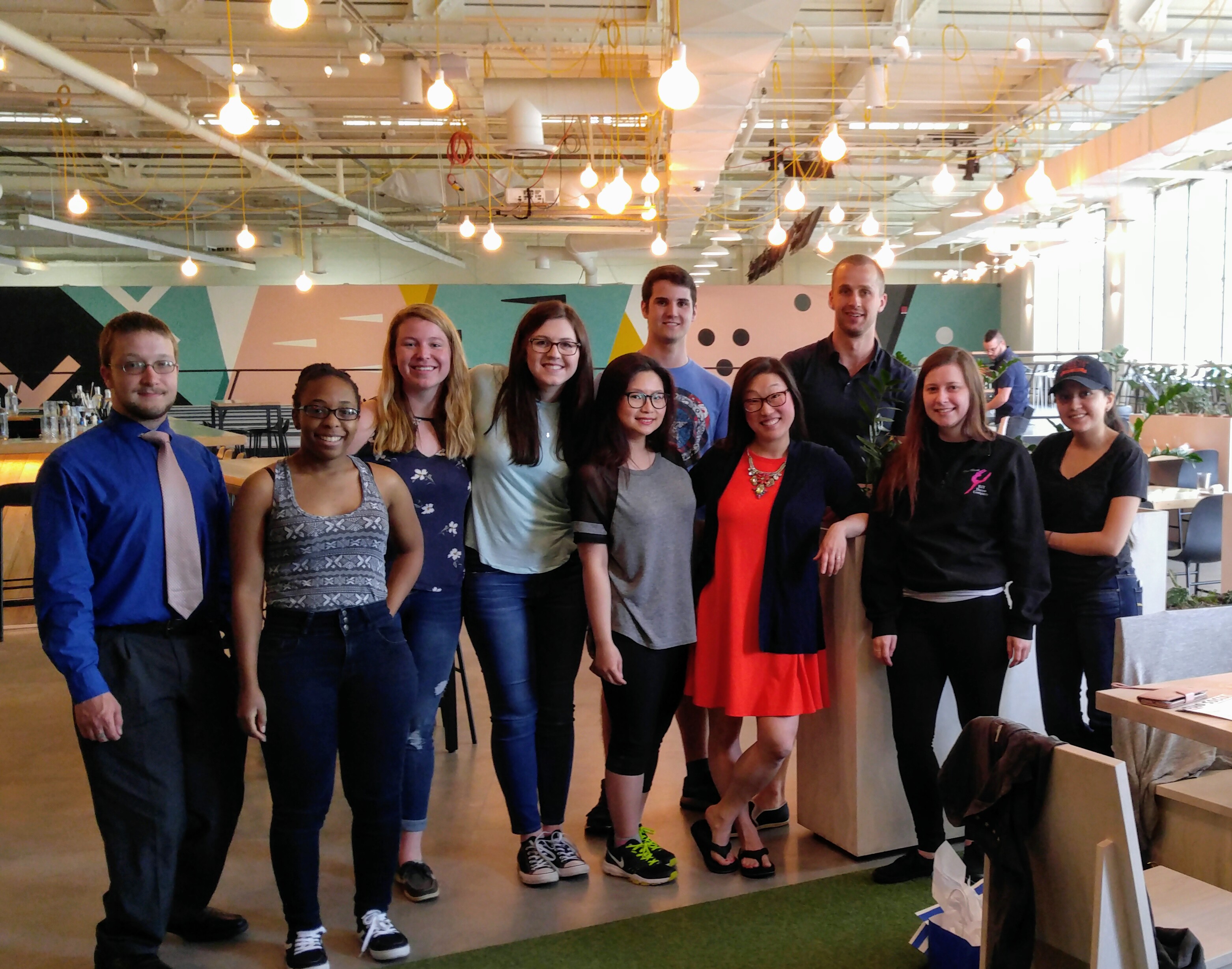

Michel Research Group at Radio Social
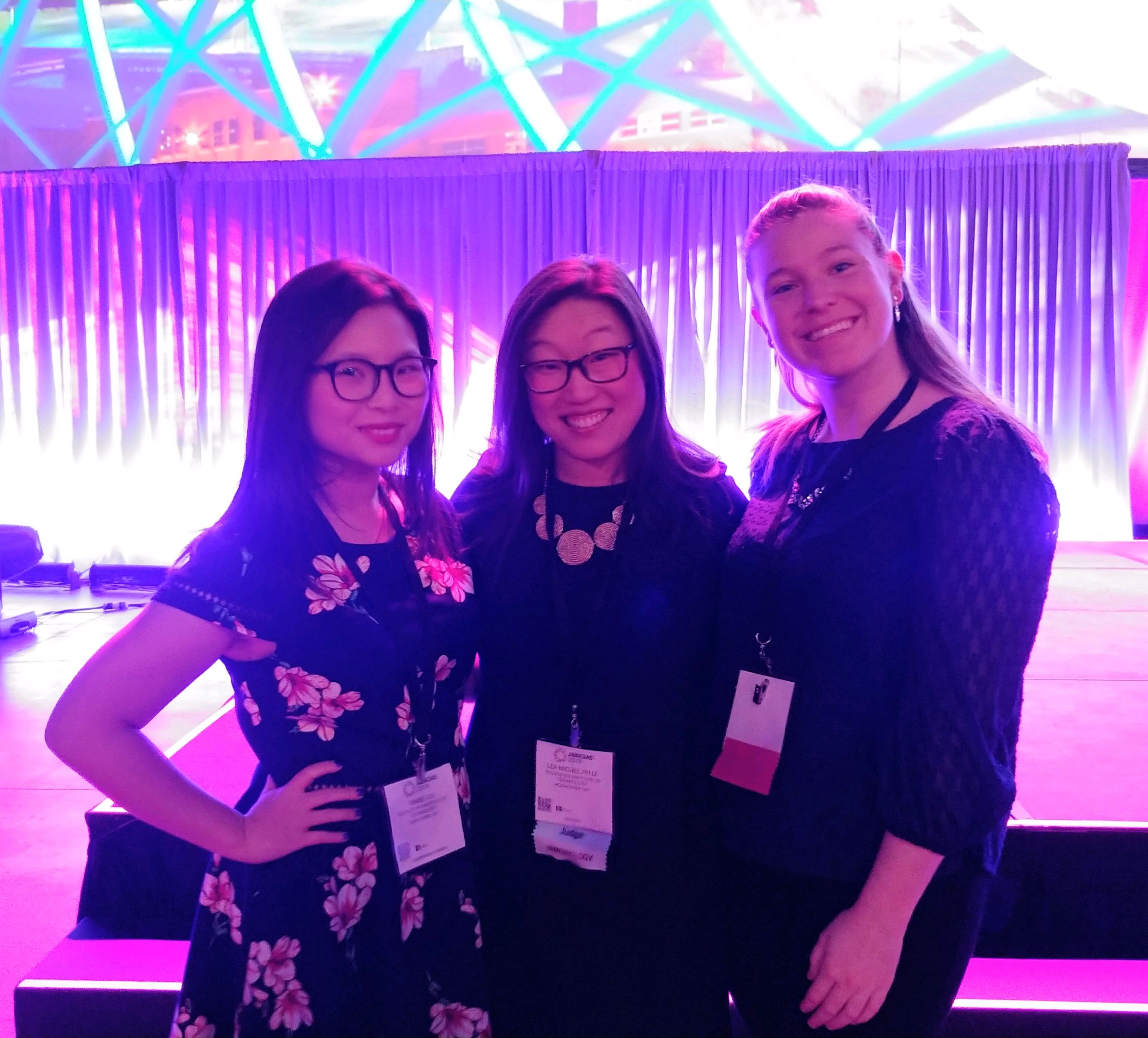

ABRCMS 2018
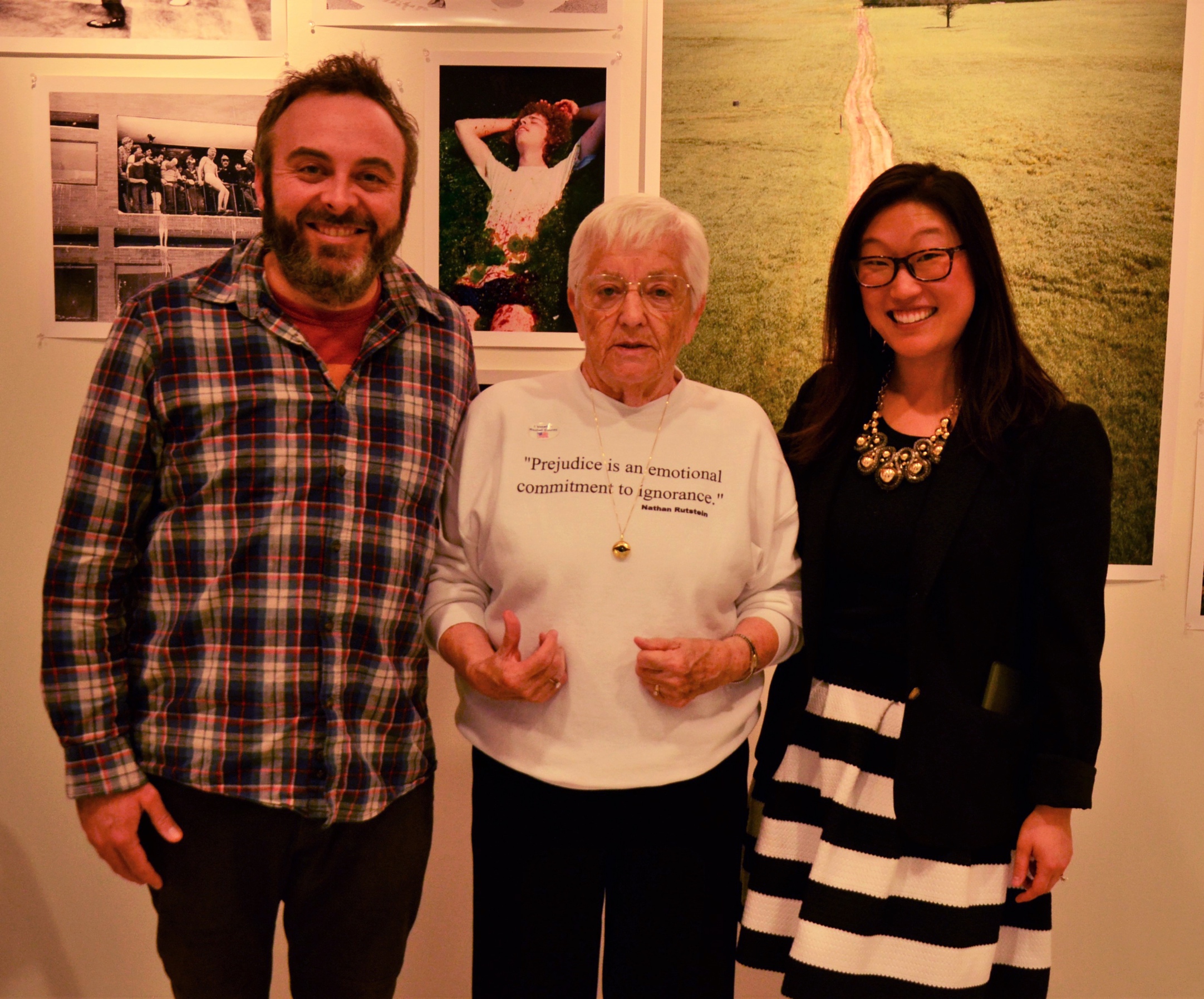

Lea and Scott Michel meet Jane Elliott!
The Michel Group takes over Dave and Buster's
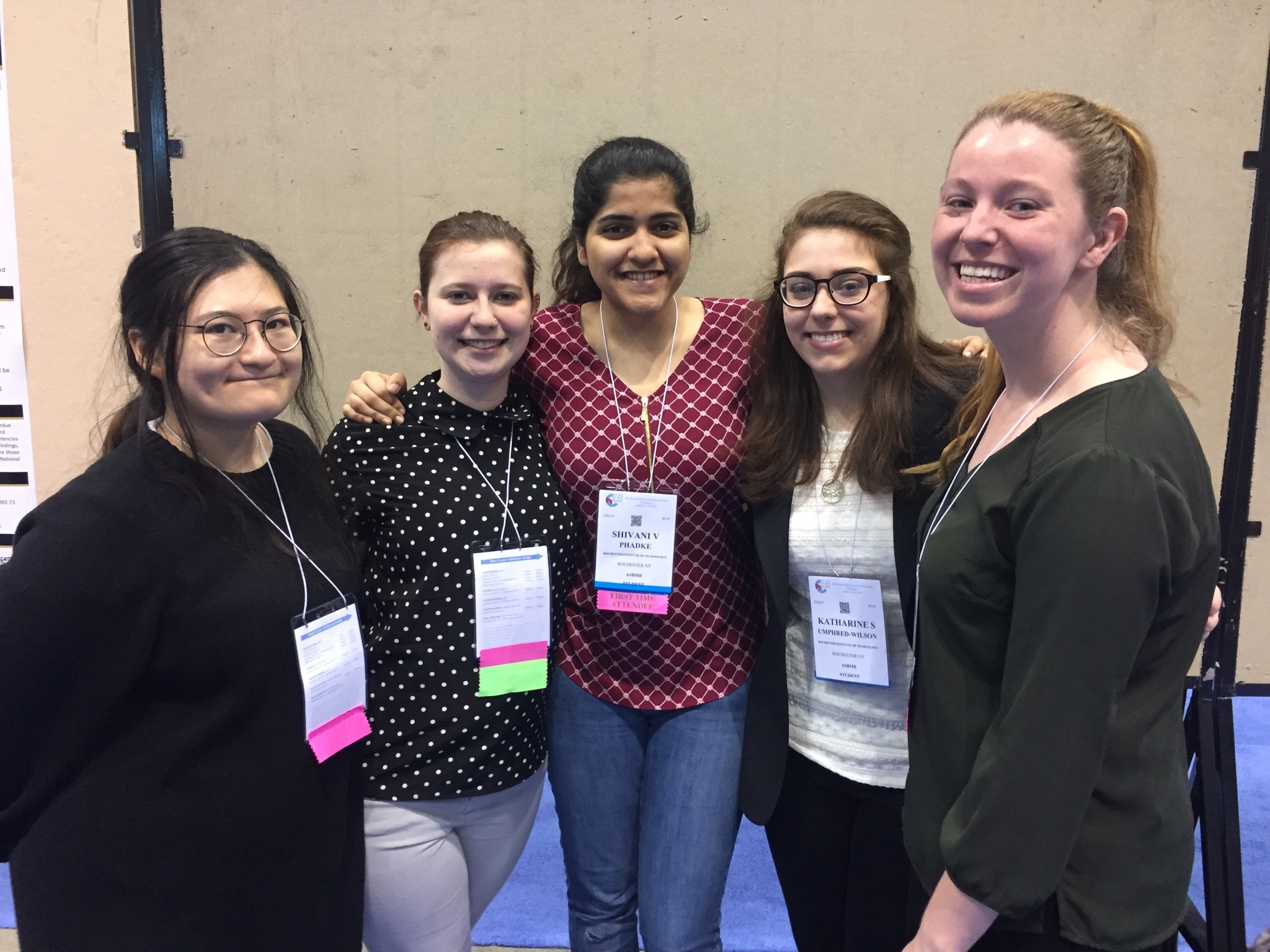

Michel Research Group at ASBMB 2017

RIT at ABRCMS 2017
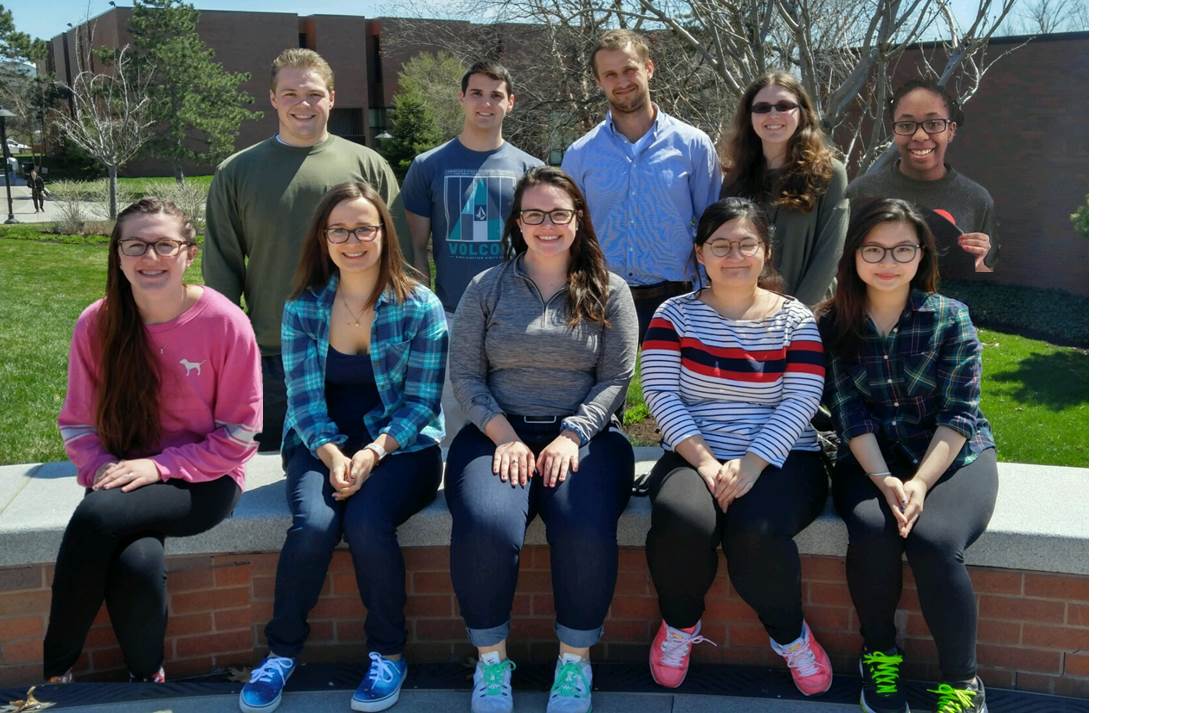

Group photo Spring 2017
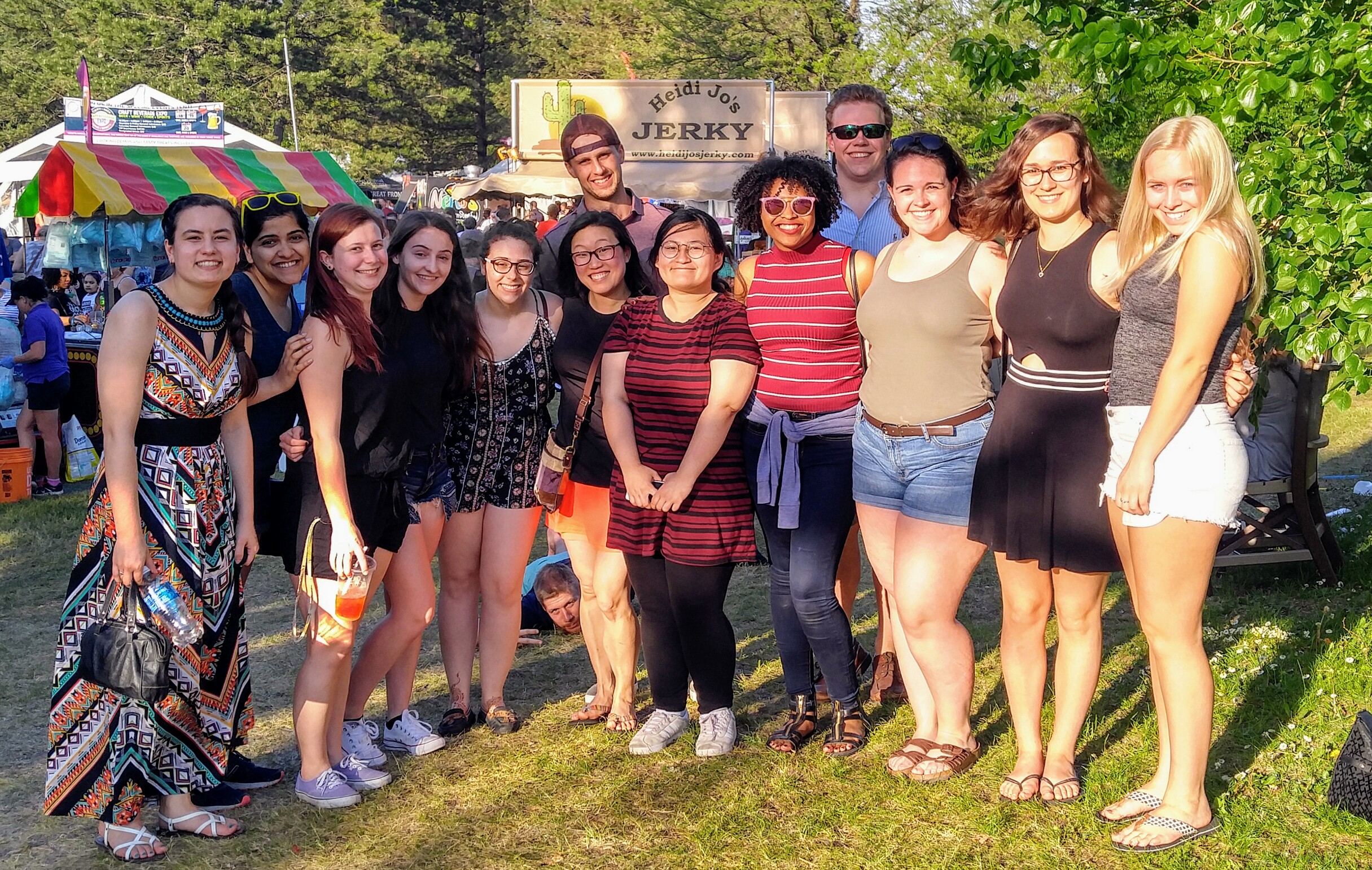

Lilac Fest 2017 (check out the photo bomber)
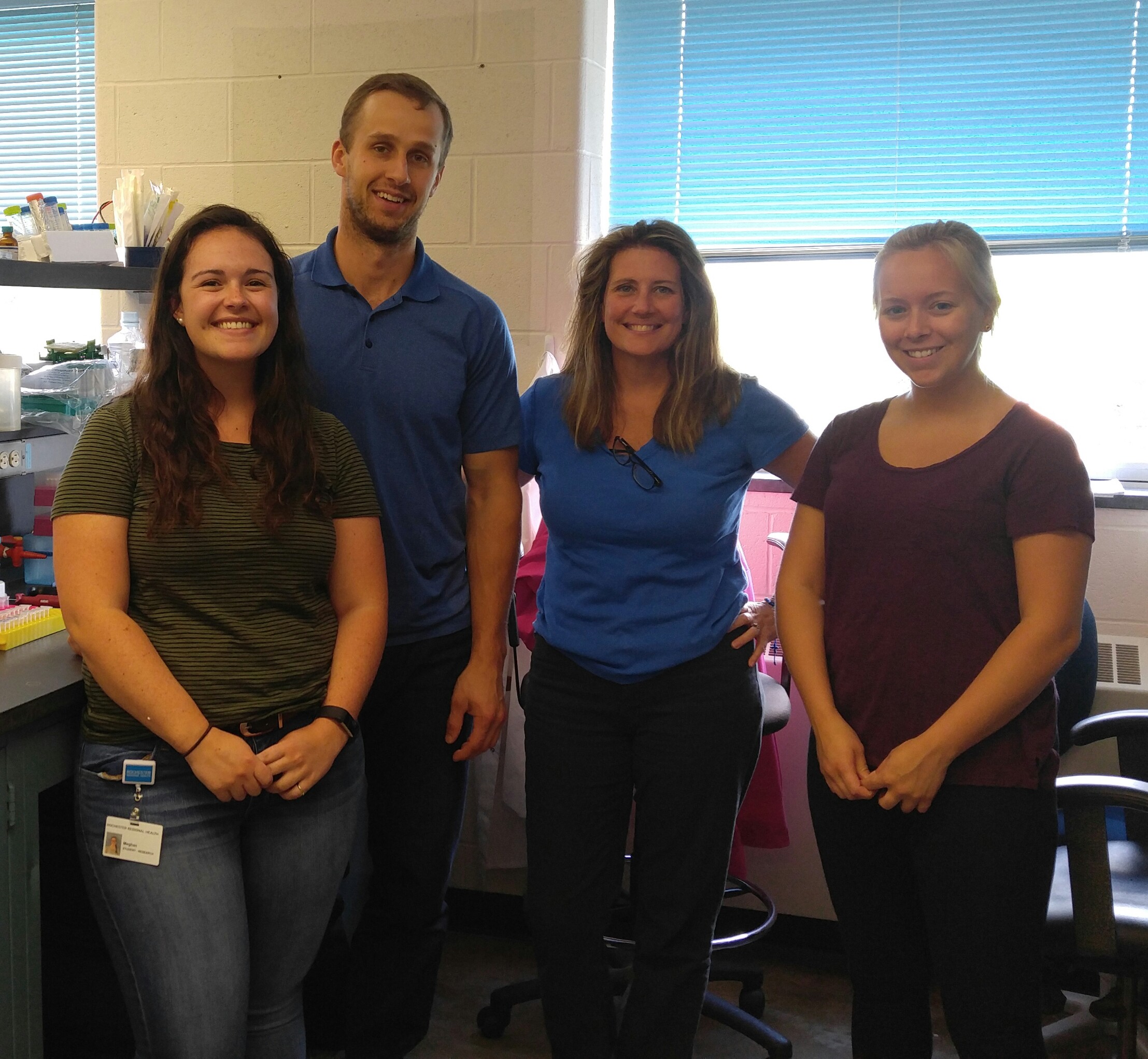

Michel group at RGH Summer 2017
Selfie outside the mouse room at RGH

Holiday Lunch 2017
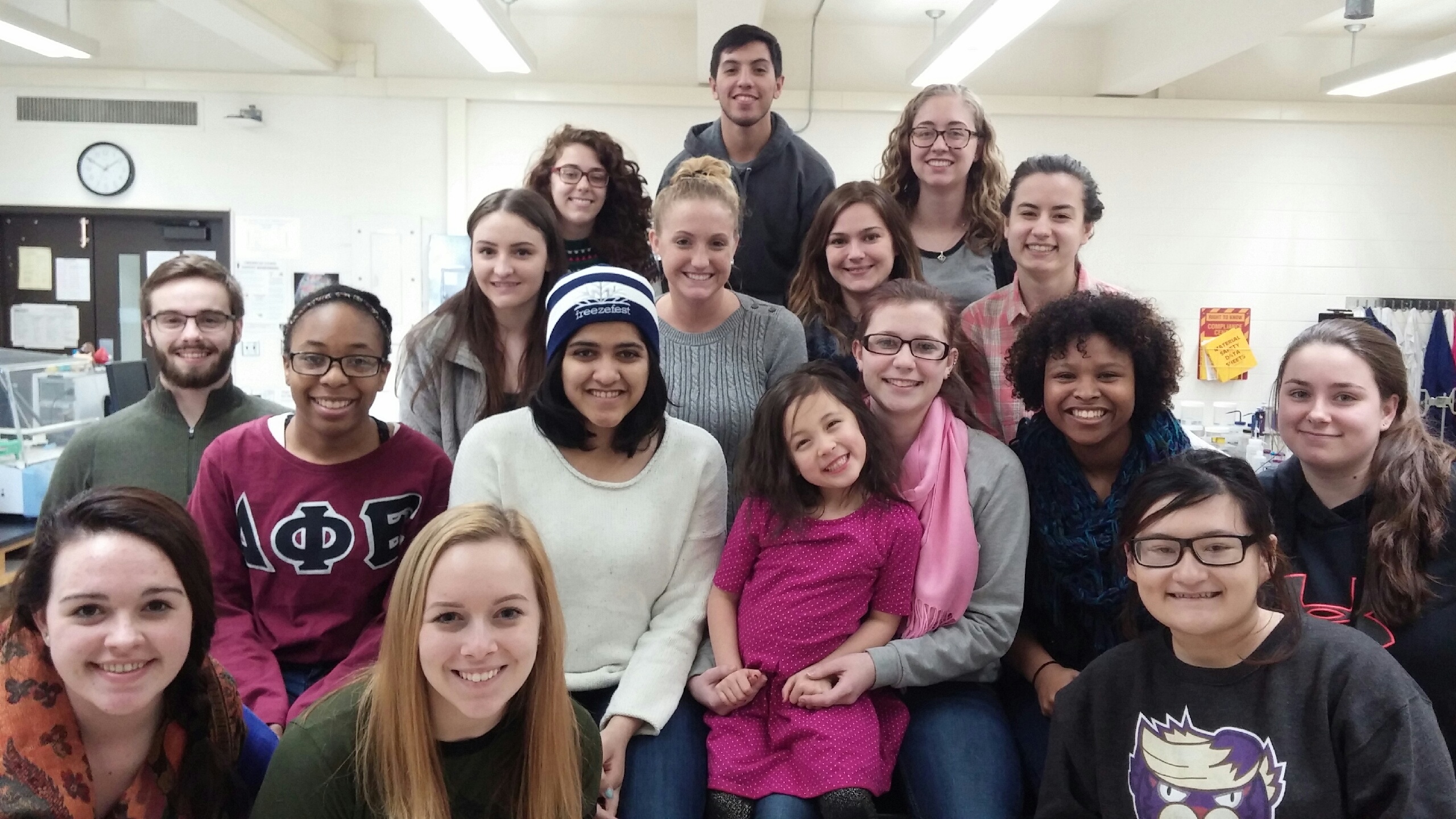

Group photo in lab 2016

ABRCMS 2016

Michel Lab Graduates 2016

Angel's grad present from Dr. Michel

Group photo 2016

Another group photo 2016

End of the year BBQ

ACS meeting in Boston

Graduates 2015

Group trip to the zoo

Summer group 2015
ASBMB 2015

ASBMB in San Diego
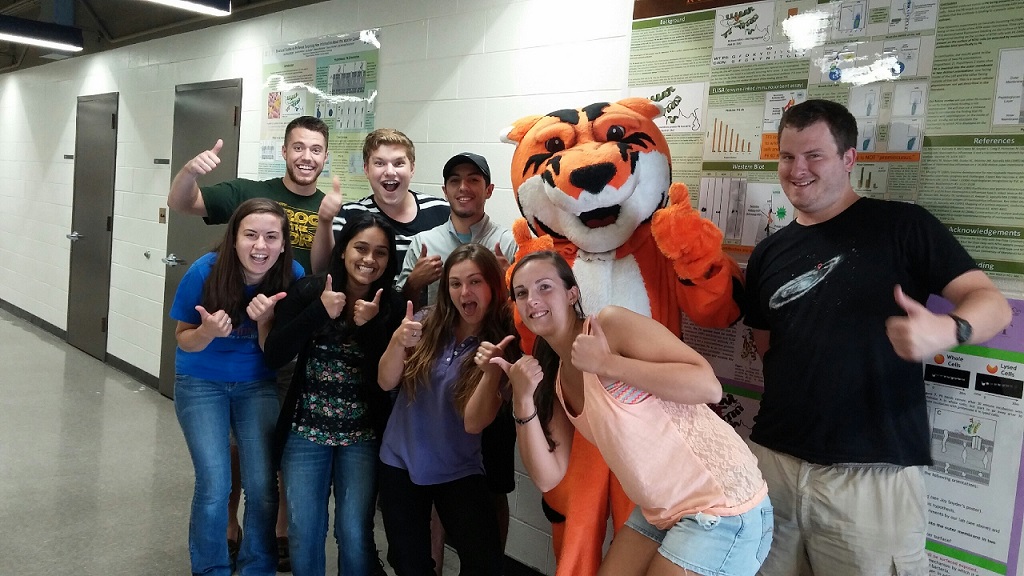
Go RIT Tigers!

Michel Group Picnic 2014

Graduation 2014

Dr Michel and Dr Tan

Graduation 2013

ASBMB 2012

ASBMB 2012

Look who we ran into at ASBMB 2012- Michel Group Alumna, Danielle (now Dr. Weekes)!

Michel Group 2012

RIT Summer Symposium 2012

Michel and Tan groups at the Zoo 2012

Graduation 2012

Just hanging with Gary Sinise in DC (ASBMB meeting 2011)

ASBMB 2011

Bowling with the Michel and Tan groups (and Autumn!)

Group photo shoot

Summer 2011

RIT at local ACS Undergraduate Research Meeting

Michel Group Summer 2010

GO BILLS!!!!

Michels go to Italy!

An updated family photo (2022)


Elsie joined the Michel family in 2022 ![]()

Scott and Lea Michel with their two children, Laralyn and Charlie

Niagara Falls 2019

Lea is a huge Gilmore Girls fan- can you spot her in this GG festival selfie?

Father's Day Gift 2019

Lea's twin sister, Meredith Vacca, recently ran a campaign to become a Monroe County Court judge. In January 2021, she became the first ever Asian American County Court Judge in Monroe County!

Laralyn and Charlie with their sweet little cousin ![]()
Archived News
Another amazing summer in the Michel Lab: Mia, Halley, Jasmine, Christina, Nikita, Mya, and Thehara

So proud of the Michel Lab 2024 Research Student Graduates: Nico, Jimmy, Lucas, Gabby, Navi, and Martina!! Off to do AMAZING things!

What a privilege to meet Dr. Rita Colwell (Distinguished University Professor at U Maryland and former Director of NSF).

Congratulations to Aidan for receiving a prestigious Barry Goldwater Scholarship!

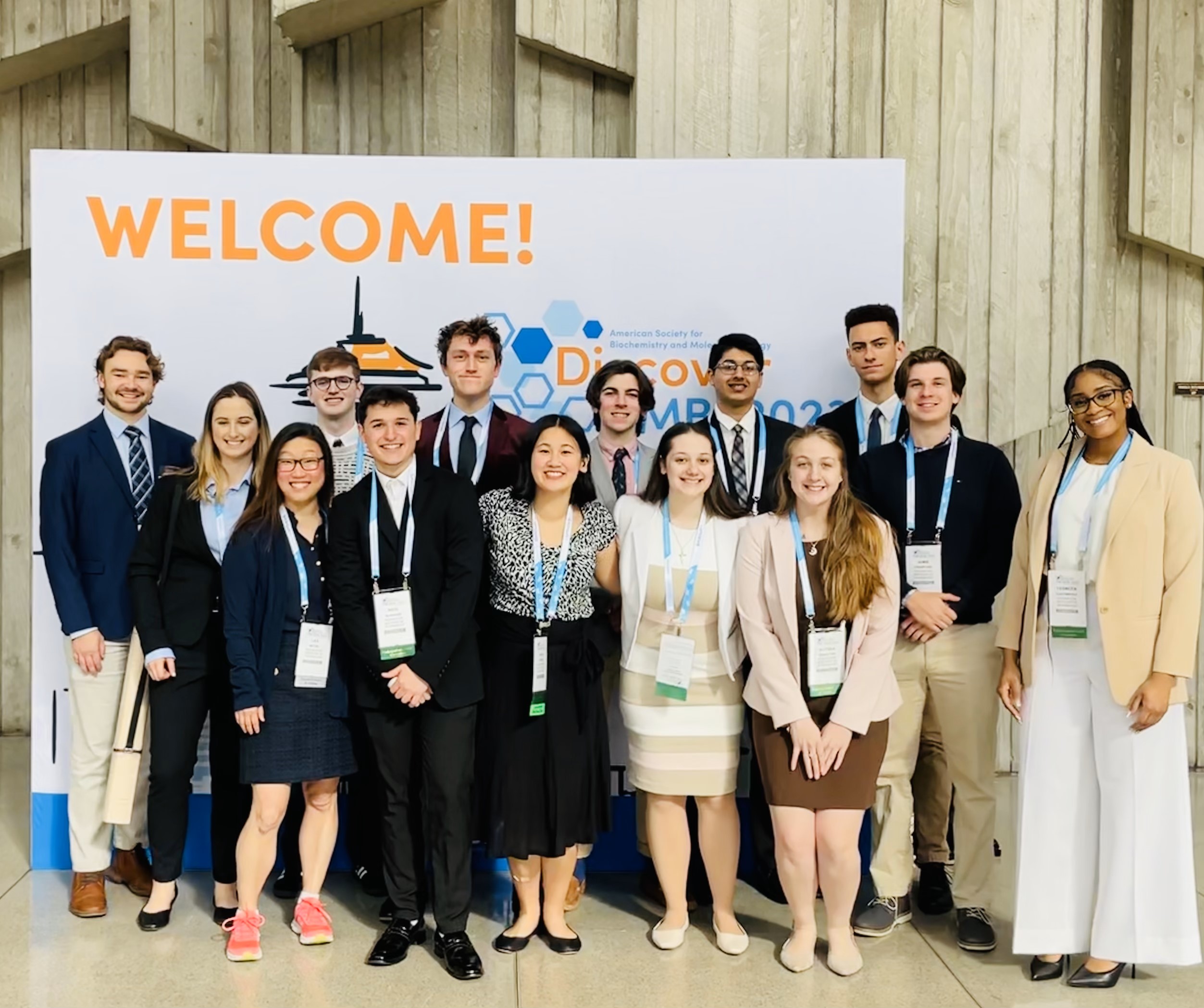
ASBMB Today featured our very own Nico Burgado and his research poster at DiscoverBMB in San Antonio!
The Michel Lab presents at DiscoverBMB in San Antonio! Congratulations, Martina, Nico, Jimmy, Navi, Aidan, Lucas, Gabby, and Tahaara! And a special congratulations to Jimmy and Navi for being inducted into the ASBMB Honor Society!
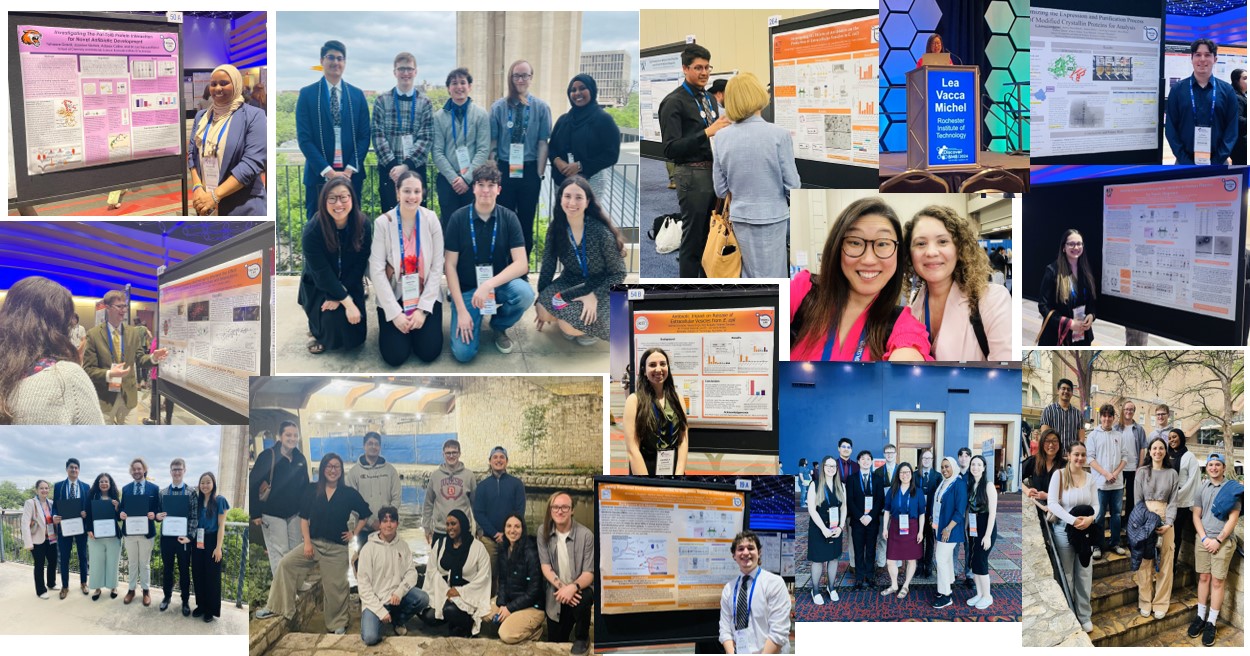

So proud of Guerline for presenting her work at the National Society of Black Engineers (NSBE) conference!

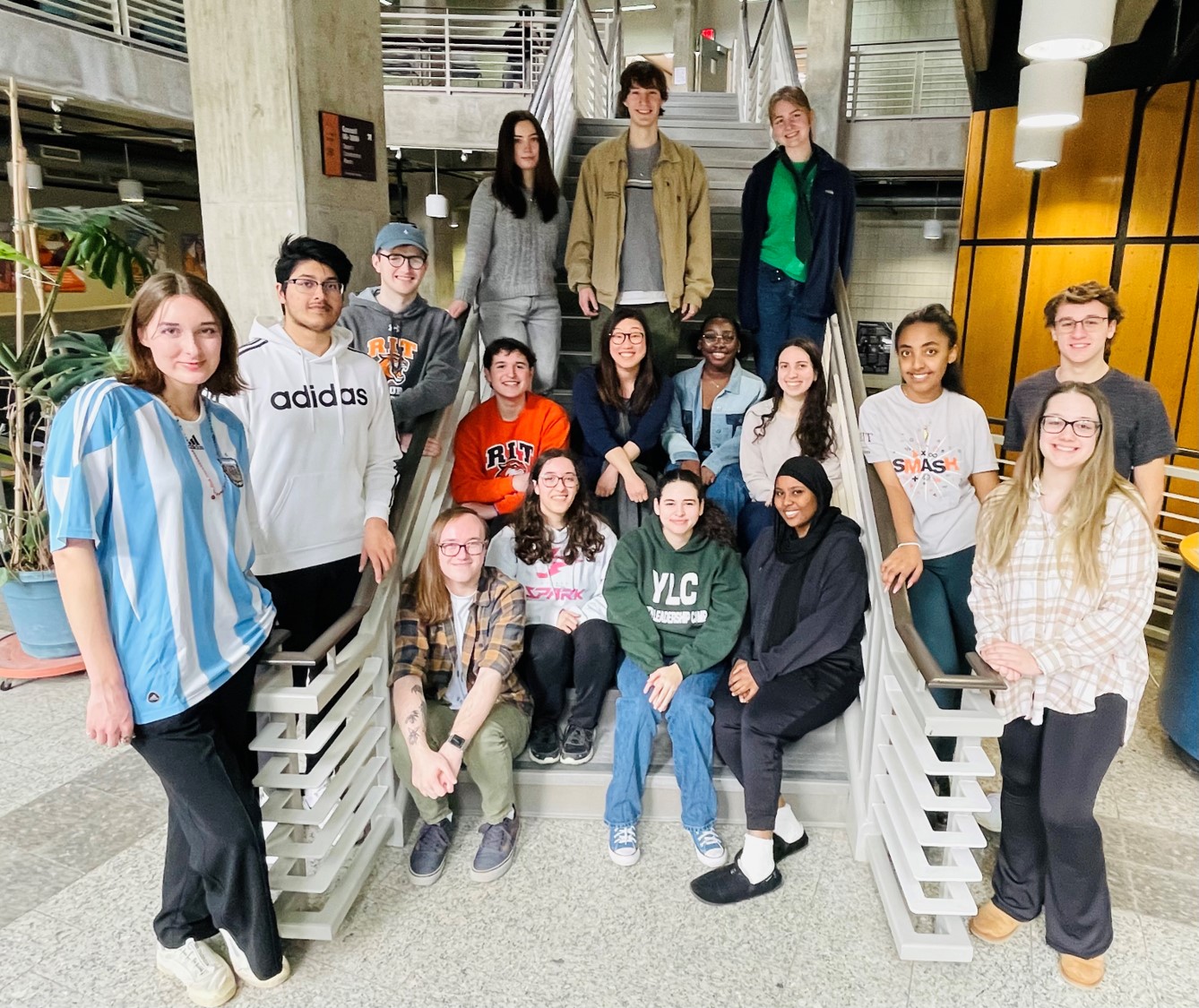
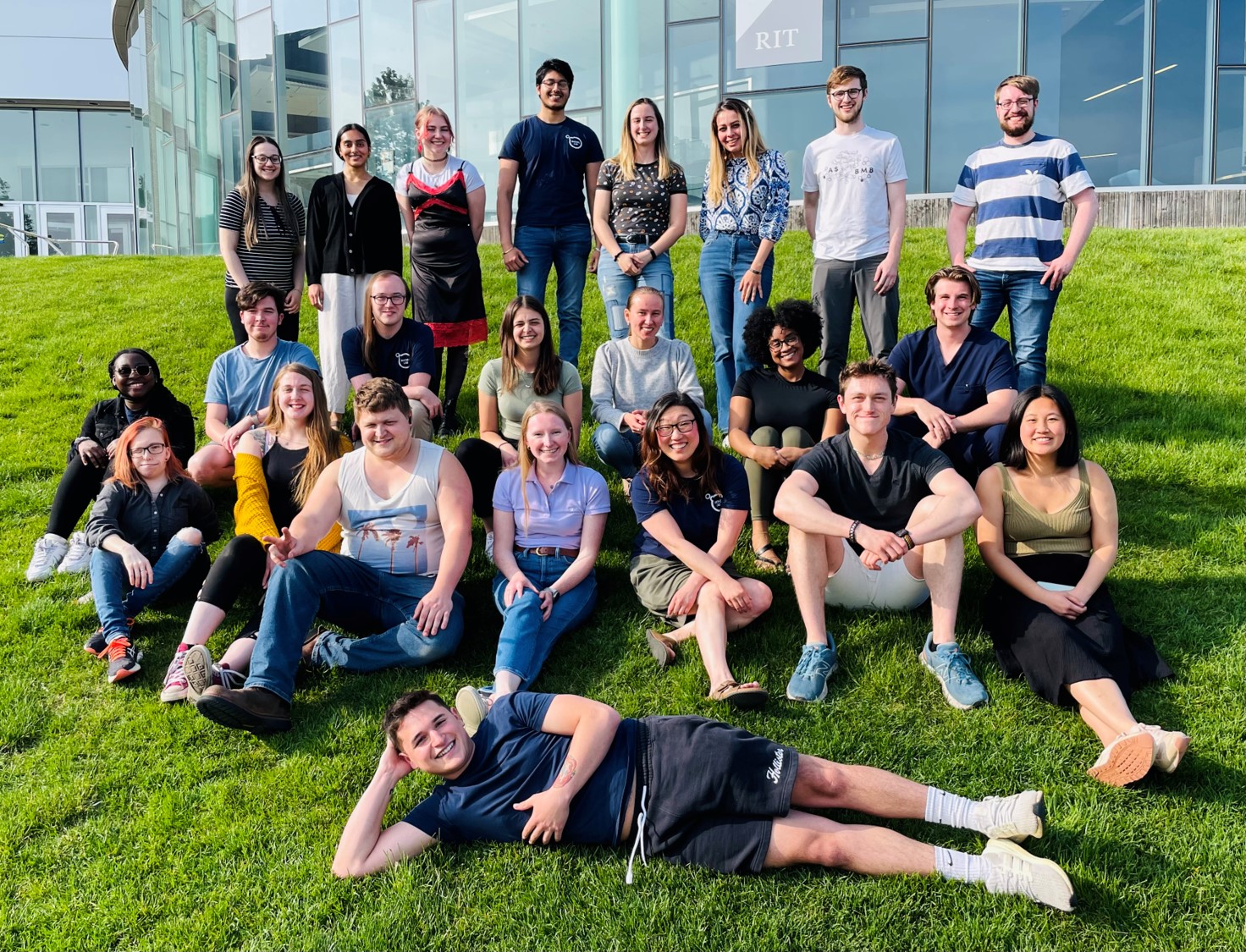
Michel Lab Spring 2024 (missing a few, but almost at full strength!)

The Michel Lab's Martina Videva was selected to be a part of ASBMB's 2023 Honor Society for undergraduate students. Check out her quote on their website!

So proud of Nico and Tahaara for representing the Michel Lab at the Annual Biomedical Research Conference for Minoritized Scientists (ABRCMS) in November 2023.

Drs. Tina Goudreau and Lea Michel published a commentary Looking at Your Lab Through a New Lens in Nature Reviews Chemistry. Check it out here! (photo featuring a few of the Michel lab members)

Summer Research 2023!

Congratulations, Anna (the COS and RIT undergraduate commencement speaker), Isabelle, Natalie, Yasmeen, Jamie, Nico, Danny, Ulysses, Alyssa, and Aoife! So proud of our Michel research group members who graduated from RIT! From med school to PhD programs at Brown and Duke to Master's programs to scientist positions at Moderna- our graduates are off to do AMAZING THINGS!
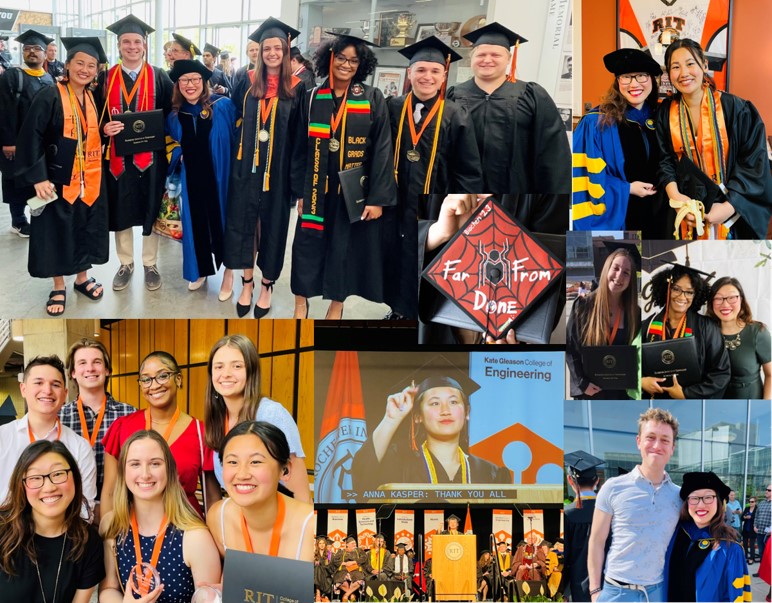

The Michel Lab celebrates the end of the school year with a potluck picnic (May 2023)

Anna Kasper won an Excellence in Student Life Award! Congratulations!

The Michel Lab and RIT was well represented at the national ASBMB meeting in Seattle (March 2023).

In March 2023, Lea had the pleasure of hosting Adam Rutherford, Geneticist and Author of How to Argue with a Racist and other amazing books, for an Inclusive Excellence seminar at RIT.

Aidan Miller and Dr. Michel had the pleasure of hosting three amazing Senior Capstone students from Rochester Prep High School over winter break. These talented young women are going places!
A huge congratulations to Isabelle for winning an Undergraduate Research Grant from the Rochester Academy of Science (rasny.org/student-grant-program)!

First Group Meeting of 2023 (we need a bigger room)!

The Michel Research Group celebrates the end of Fall semester 2022 (did I mention this is only 2/3rds of the group)!

Lea Michel is named the first RIT College-level Director of Diversity, Equity, and Inclusion.

Michel lab alumni were featured in Lea's ASBMB TODAY essay about inclusive social events, "All alone in a crowd" (August 2022).

It's was a busy summer in the Michel Lab. Very proud of what these students learned and accomplished! Left to right: Martina, Navi, Lea, Jimmy, Isabelle, Yasmeen, Alyssa, Jasmine, Aidan, Jada, and Olivia

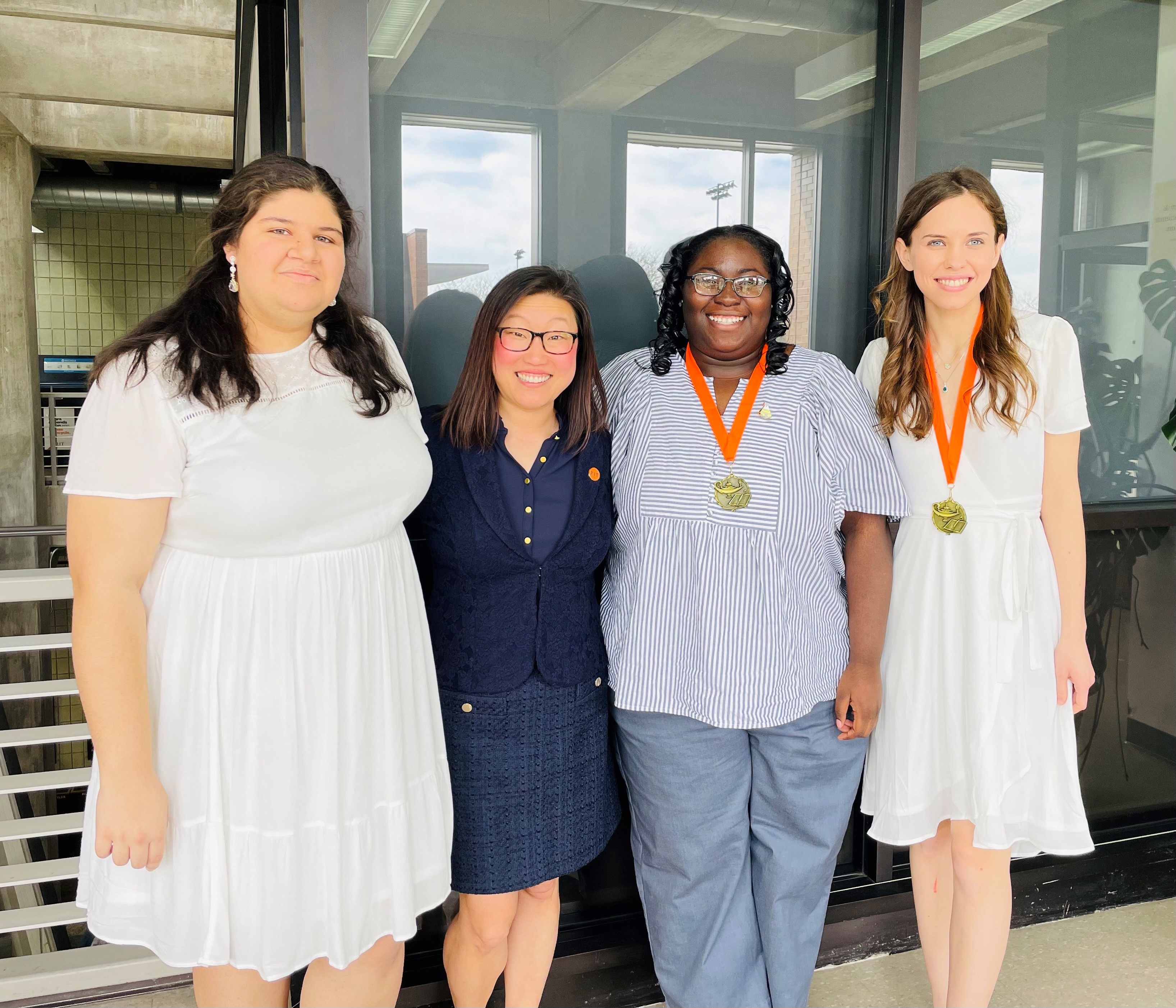
So proud of our Michel Research Student Graduates, Class of 2022! Congratulations, Janai, Zach, Katie, Milena, and Jonathan!
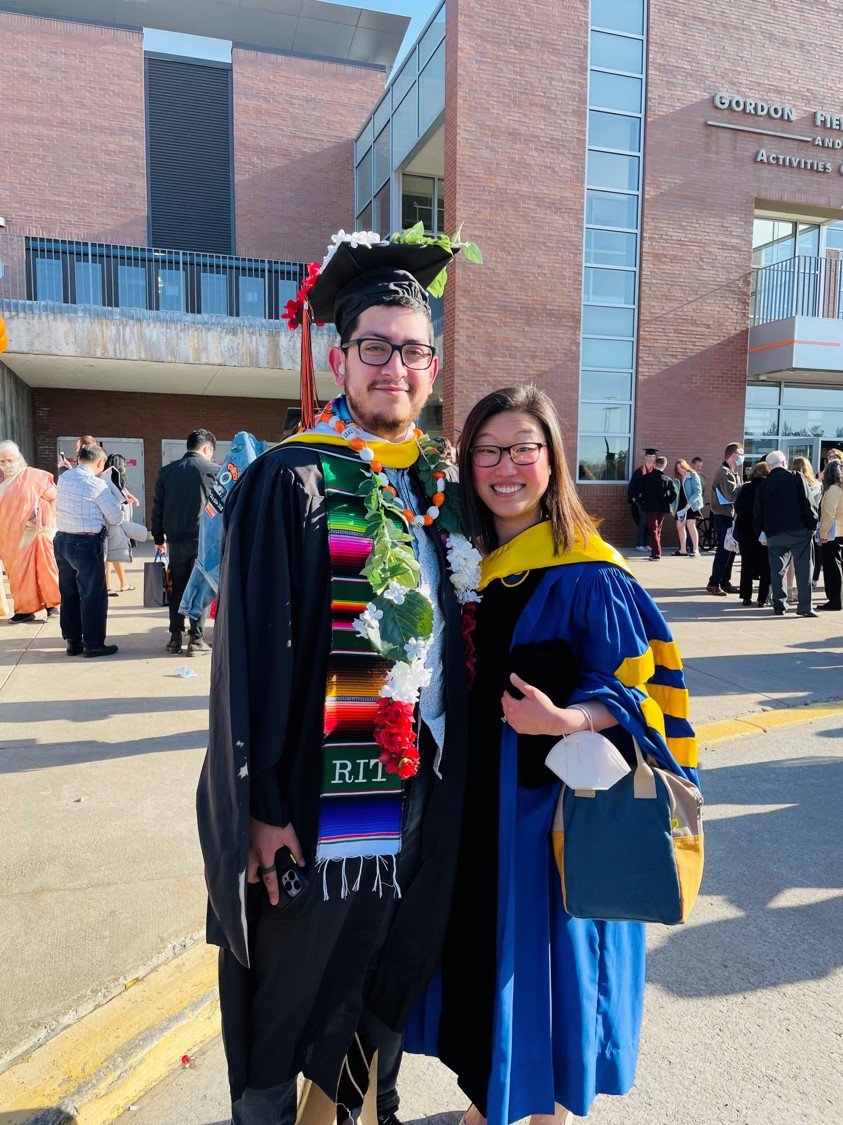
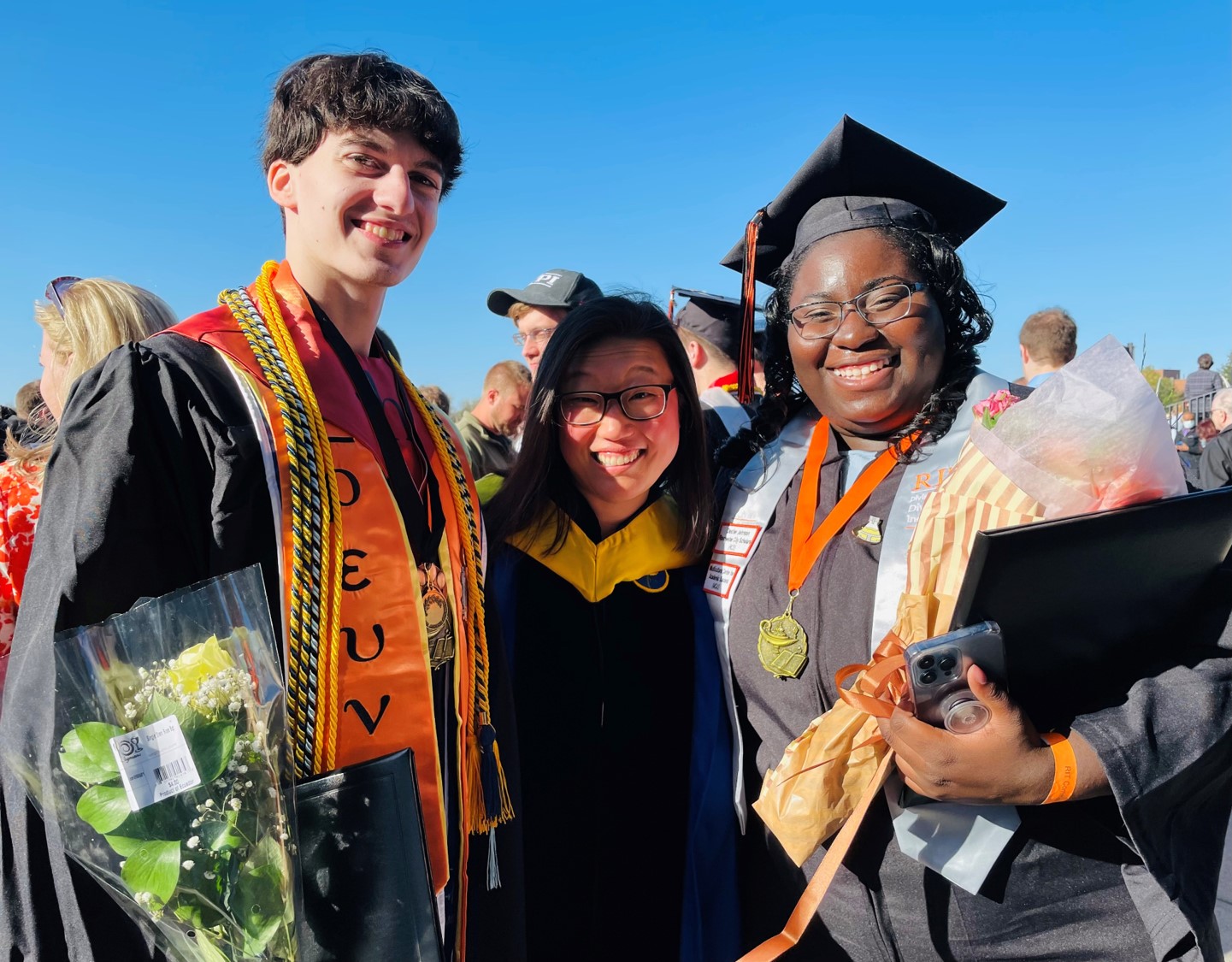



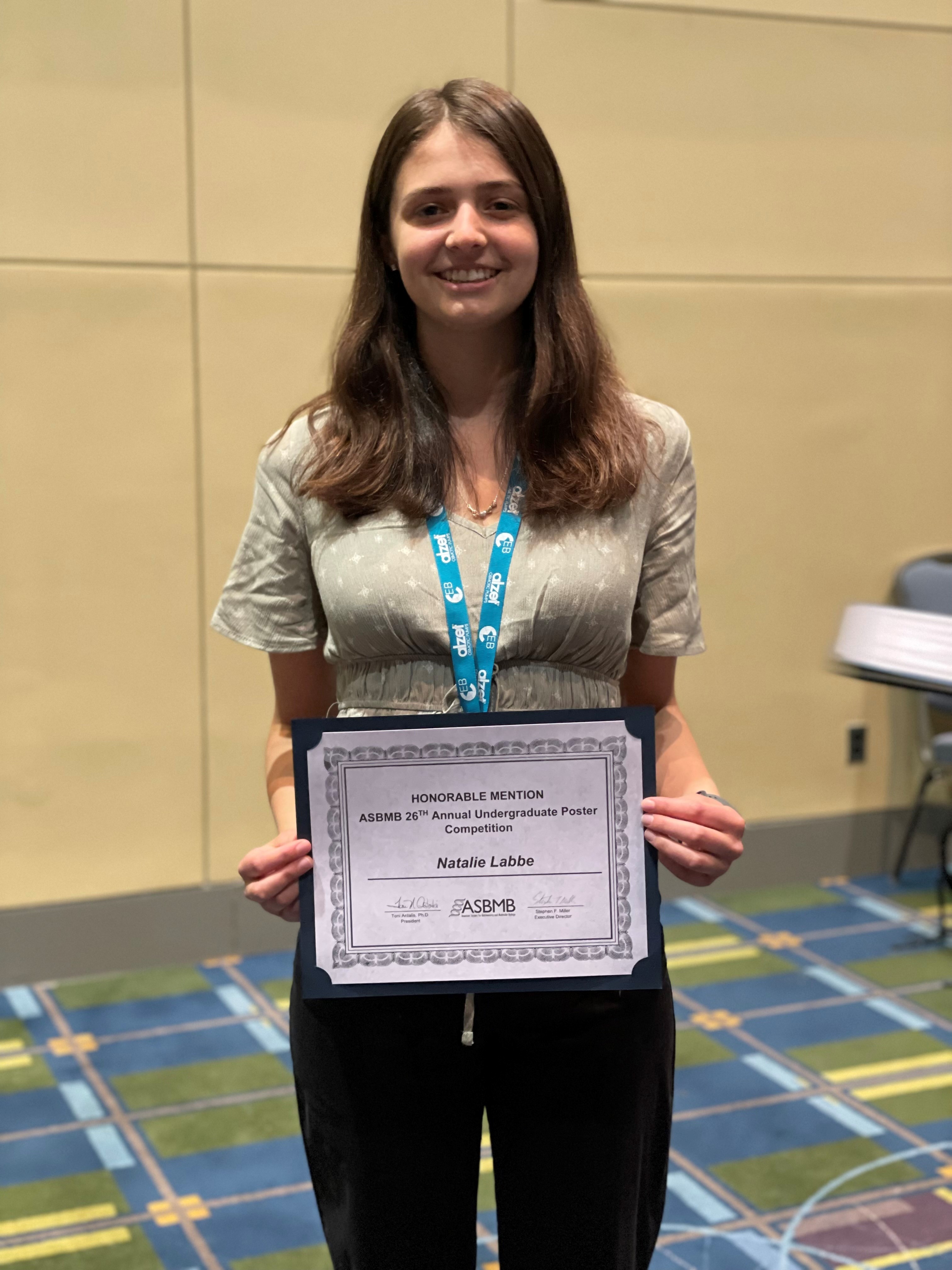
Eight students from the Michel Group presented their research at the 2022 ASBMB Annual Meeting in Philadelphia, and Natalie Labbe won a Poster Award at the Undergraduate Poster Competition! (Left to right, Back row: Zach Williams, Janai Perdue, Natalie Labbe, Grace McGinnity, Front row: Isabelle Pilo, Anna Kasper, Lea Michel, Katie O'Neill-Knasick, Milena Dinu)
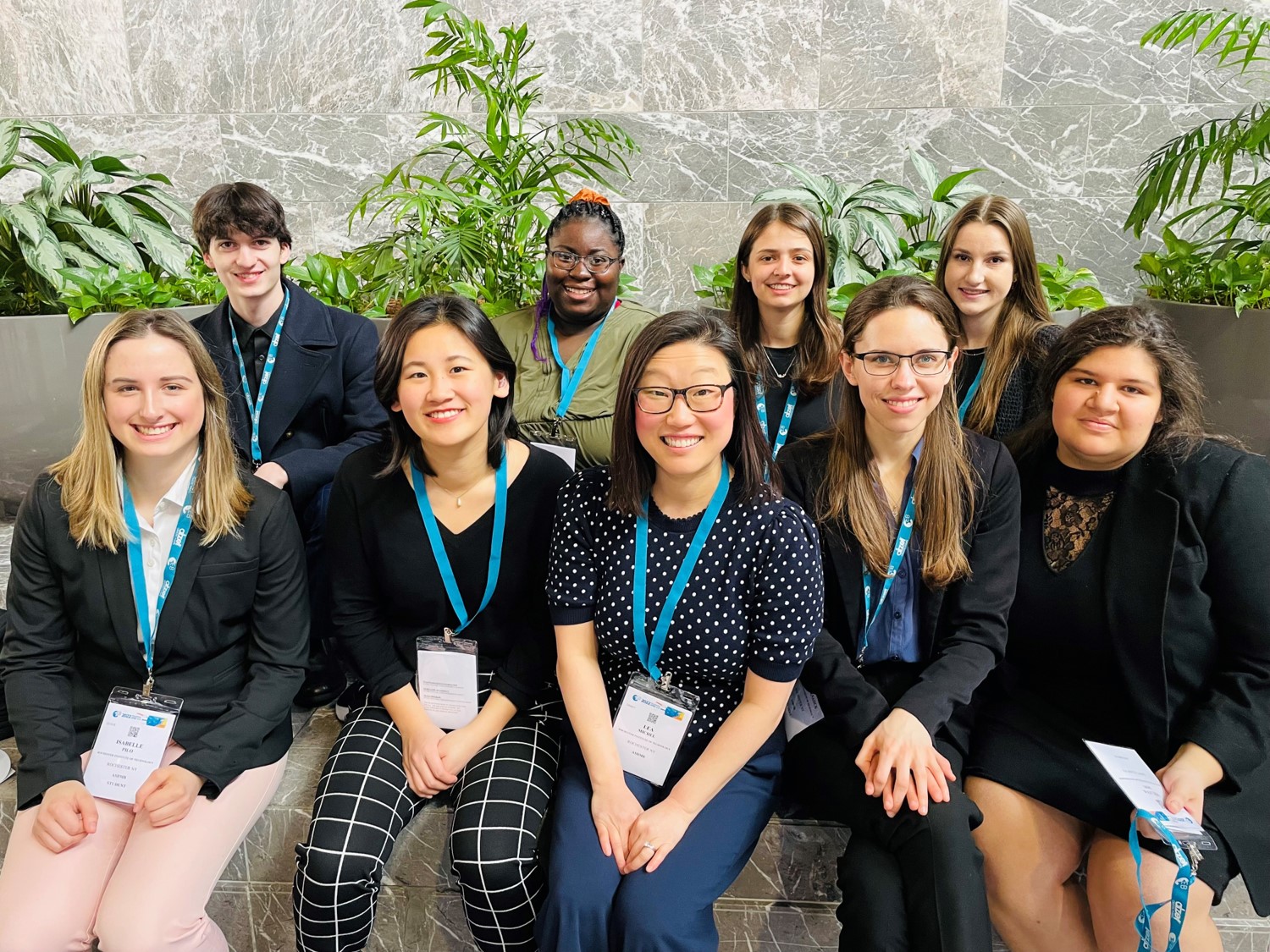


Lea Michel and Dr. Desiree Forsythe (RIT's Inclusive Excellence Program Director) wrote an article on uncompensated labor that was featured in the ASBMB Today magazine in March 2022.
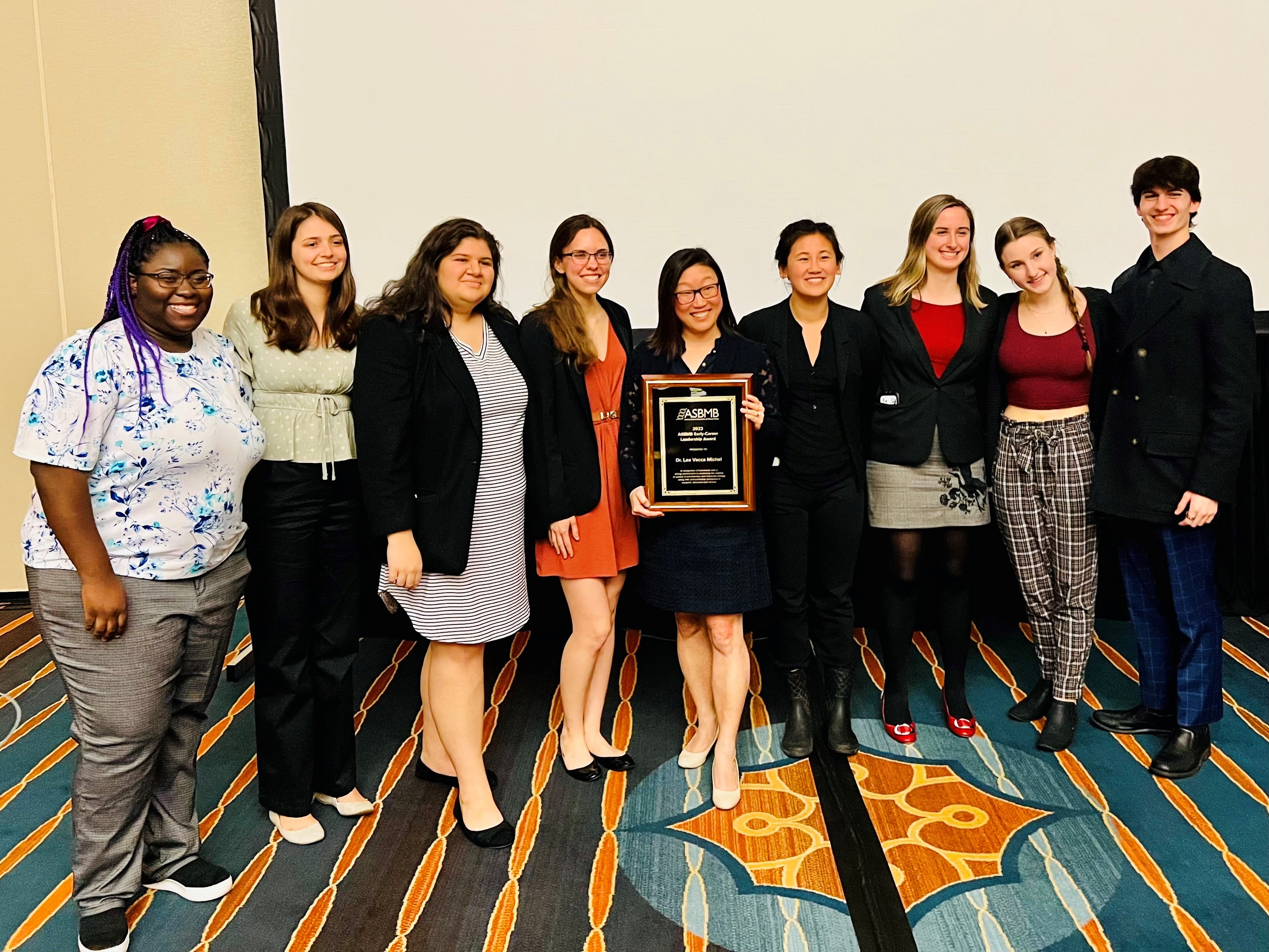
Lea Michel was named the winner of the 2022 ASBMB Early-Career Leadership Award! The award was established by the ASBMB Women in Biochemistry and Molecular Biology Committee to recognize individuals with a strong commitment to advancing the careers of women in biochemistry and molecular biology along with demonstrated excellence in research, discovery, and/or service. ASBMB Today article

Congratulations to our very own Natalie Labbe for earning a Marion B. Sewer Distinguished Scholarship for Undergraduates Award from the American Society for Biochemistry and Molecular Biology!

Congratulations to Tyler Pugeda for being selected for a prestigious Fulbright fellowship to study human brains afflicted with Alzheimer's disease in Germany.