Student to Student: Brittle stars
By Alexandria Shumway
Alexandria Shumway is a fifth year (graduate) bioinformatics student who is currently working on her masters. Her mentor/advisor is Dr. Hyla Sweet.

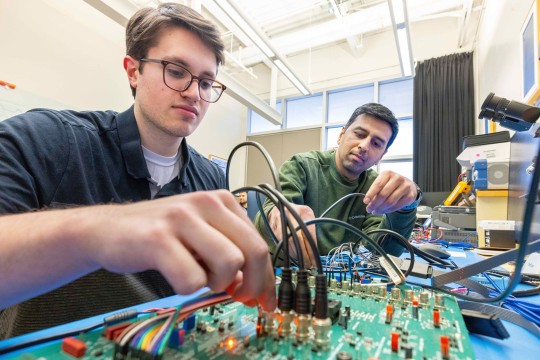
Currently, my research focuses on the development of the Ophioplocus esmarki brittle star. Brittle stars belong to the phylum Echinodermata, which contains over 7,000 living species of marine organisms. Echinoderms are unique in their five-fold radial symmetry, ability to reproduce asexually, and their utilization of a water vascular system. Within this phylum, the Ophiuroidea class contains both species of brittle star and basket star, with the O. esmarki brittle star being the focus of my research. Echinoderms have two independent nervous systems: a bilateral nervous system that controls the larva early in life and a second nervous system with five-fold symmetry that forms later to control the juvenile and adult body systems. This project focuses on the evolution of neural development between these two modes.
This research is part of my ongoing Master’s thesis, which focuses on creating and analyzing transcriptomes from two different stages of development and identifying transcripts involved in the development of the nervous system in O. esmarki.
In the Spring of 2018, we sent out samples from larval and juvenile stages of development for sequencing and assembly at the University of Rochester Genomics Center. Once the transcriptomes were complete, we were able to identify different molecular and biological roles. By comparing these results between the two stages of development, any difference can show the changes in RNA expression. We are currently looking at different neural transcripts originally identified in the model sea urchin, Strongylocentrotus purpuratus. After locating these transcripts in our brittle star transcriptomes, we can validate findings through phylogenetic tree analysis giving us stronger confidence in transcript identity.
Ultimately, we will look at the spatial expression patterns of these neural transcripts to better understand the development and transition of the nervous system from bilateral to five-fold symmetry. This species O. esmarki is an important organism to study, due to its two modes of contrasting development. By creating the first transcriptome for this species, the data it provides will serve as a foundational tool for future studies. Information gathered by this project should help to strengthen the knowledge about brittle star development and provide a substantial resource to fuel other projects for future RIT biology students.
How did you come to study bioinformatics at RIT?
Before RIT, I had never heard of bioinformatics before. I knew I was interested in the realm of science and decided to branch out into this new major and see if it was a good fit. By my second year, I slowly became more comfortable with my major-specific classes and knew this was the right path for my future.
You are researching brittle stars. How did you become interested in this topic?
In the Introductory Biology II class during my first year, I was placed in a group to study the effect of wound type on regeneration abilities in the Ophioplocus esmarki brittle star. We were able to pick from a list of group projects ranging from cancer imaging to soil organisms, to organic insecticides. However, I had never heard of this particular organism and it instantly piqued my interest. I joined Dr. Sweet’s research group soon after the class ended and became more interested in developmental and bioinformatics-based research.
Any other research related experience you would like to share?
In the summer of 2018, I applied to over 10 REUs along with a few other summer laboratory positions. With one rejection followed by another, I began to question my ability to succeed in a scientific field. However, these failures helped me to recognize and improve my weak spots. I realized I needed to grow and become more fluent in the ‘wet lab’ techniques that were not as prevalent in my bioinformatics degree. That Fall, I was able to obtain my first publication and present at multiple scientific conventions. I enrolled in a Genetic Engineering course and slowly improved my comfort level and expanded my skills with each passing day in the lab. With these opportunities, I then became a peer mentor and vocalized my struggles in the scientific field to place importance on failure and self-analysis to enable one’s career growth.
Have you studied abroad? If so, what advice would you like to share with other students?
I was lucky enough to do research abroad this past summer in Kiel, Germany. Taking off to a foreign country, with cities I had never visited, a language I could not speak, and an entire professional scientific community that I had never encountered, was the biggest risk I had ever taken both personally and professionally. I was sponsored by the DAAD RISE program and worked on intestinal inflammation under a Ph.D. student. This topic was really important to me because my grandmother and best friend are affected by inflammatory bowel disease. This experience was able to reaffirm my beliefs to pursue a Ph.D. and helped to assert my goals for the future.
My advice to other students would be to take risks. I never thought I would be accepted to this competitive program or have the guts to go so far out of my comfort zone, but it paid off in unimaginable ways.