Engineering faculty member receives NIH grant to develop biotechnology to better detect sepsis
New nanodevice could detect drug-resistant bacteria faster and more accurately, increasing chances of treatment and recovery
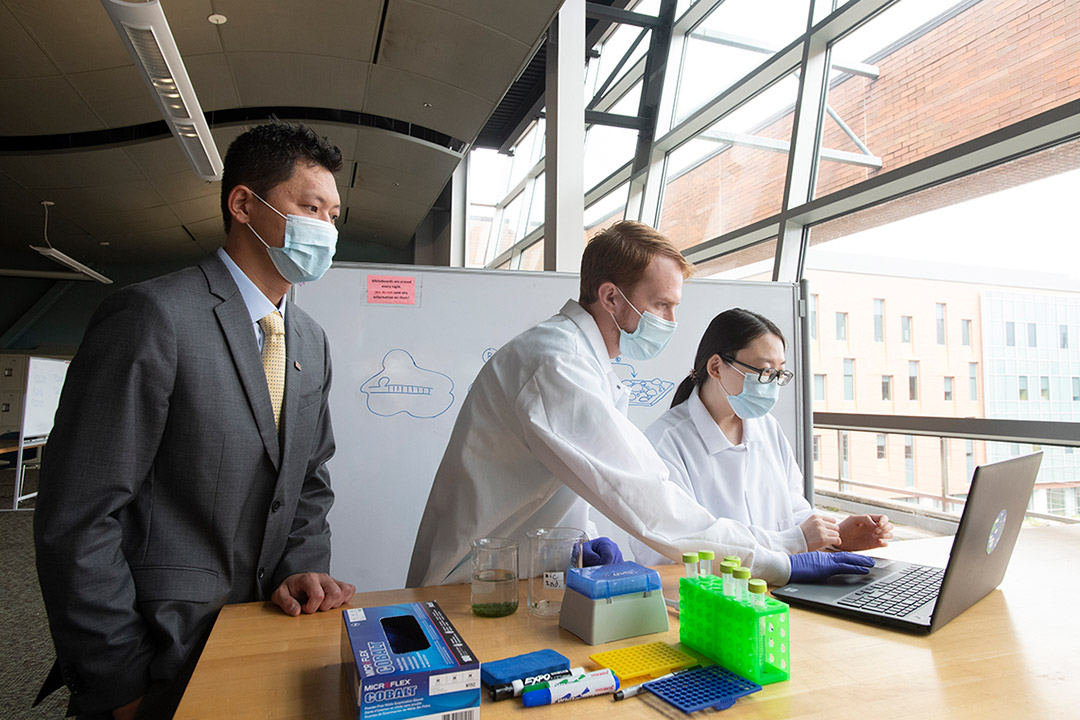
A. Sue Weisler
Ke Du, assistant professor of mechanical engineering, left, works with Jacob Waitus, center, a BS/MS chemical engineering and materials science student, and microsystems engineering doctoral candidate Li Liu in Du's Nano-biosensing, Nano-manufacturing and Nano-materials Lab.
As one of the leading causes of death in hospitals, sepsis becomes more complicated with the rise in bacteria most resistant to some of today’s antibiotics. If physicians can detect onset earlier, treatments could begin sooner to improve patient mortality.
Ke Du, a mechanical engineering faculty-researcher at Rochester Institute of Technology, will be developing a microfluidic device to improve detection of drug resistant bacteria in blood, one of the most common pathogens causing sepsis.
Du recently received a National Institutes of Health (NIH) grant to develop a detection system and to further clinical studies of how the bacteria can be targeted for intervention. The five-year, $1.8 million award is part of the NIH’s Maximizing Investigators’ Research Award for Early State Investigators program. This is given to faculty-researchers who have demonstrated foundational biomedical technologies, such as prototype devices and applications, that could advance important medical breakthroughs. One of the many areas under review by the NIH is sepsis—specifically toward effective treatment options, better diagnosis technologies and to further clarify key risk factors.
“Even one small amount of bacteria can cause this severe infection. How can you find it? Then how do you know that this bacteria is the drug-resistant bacteria? That is why we are working on the CRISPR assay—the gene editing method to specifically target drug resistant bacteria strains. The idea is, if you can detect the drug resistant strains as early as possible, then the doctors would know which antibiotics or treatments can save lives,” said Du, an assistant professor of mechanical engineering in RIT’s Kate Gleason College of Engineering.
Over the first phase of the project that started this summer, Du and his project team will integrate several sensor instruments and genotyping protocols to assess the genetic information to find specific drug-resistant strains. These studies will be the basis for an optofluidic sensing device that can directly diagnose sepsis in animal models.
“Detection needs to be accurate; this means using cell culturing,” Du explained. “Using a centrifuge and other purifying processes, you can separate the bacteria from bodily fluids and let them grow in culture media. But this takes several days and requires even longer time to test antibiotic susceptibility. This may set a person back for the most effective treatment. That is one of the things we are trying to improve with this new device.”
According to the National Institute of General Medical Sciences, nearly 1.7 million adults in the U.S. develop sepsis, and a quarter of that number die from the infection. Some infections cannot be resolved by general antibiotics due to strains of bacteria mutating making it a challenge to provide adequate treatment.
Since he began at RIT, Du has built an interdisciplinary research team, including undergraduate and graduate students, in his Nano-biosensing, Nano-manufacturing and Nano-materials Lab (N3). He is also part of RIT’s Personalized Healthcare Technology initiative that aims to bridge engineering and healthcare. His work involves improvements to biosensing devices using new materials and nanomanufacturing functions as options to distinguish hard-to-differentiate bioparticles. These technologies can advance point-of-care and rapid detection technologies needed today.
“This is important work, and I have found many talented undergraduate and graduate students interested in this,” he said. “We are trying to use a very powerful genetic method to detect the drug-resistant strains. This will be a trending project and we hope all these pieces will come together.”