Bioinformatics Master of Science Degree

Bioinformatics
Master of Science Degree
- RIT /
- Rochester Institute of Technology /
- Academics /
- Bioinformatics MS
A bioinformatics master’s degree prepares you to tackle complex problems in biology using big data, data mining, machine learning and modeling.
30%
Merit Scholarship
$1M+
Equipment in Genomics Lab
Overview for Bioinformatics MS
Why Pursue an MS in Bioinformatics at RIT?
STEM-OPT Visa Eligible: The STEM Optional Practical Training (OPT) program allows full-time, on-campus international students on an F-1 student visa to stay and work in the U.S. for up to three years after graduation.
Jobs at Industry Leading Companies: Graduates from our bioinformatics program have secured positions at esteemed institutions such as Pacific Northwest National Laboratory, Personal Genome Diagnostics, University of Rochester Genomics Research Center, and Asuragen, Inc., highlighting the program’s efficacy in facilitating career advancement.
Educational Enhancement: Our bridge program supplements your existing education, ensuring a seamless transition into the rigorous curriculum of bioinformatics.
Tailored Learning Experience: Delve into a customized curriculum offering a robust blend of biotech and computer programming, providing you with a solid foundation in both fields.
Cutting-Edge Research: Engage with faculty actively involved in groundbreaking research spanning molecular evolution, ecological modeling, cancer chromatin, machine learning, genomics, and forensic science, enriching your academic journey with the latest advancements in the field.
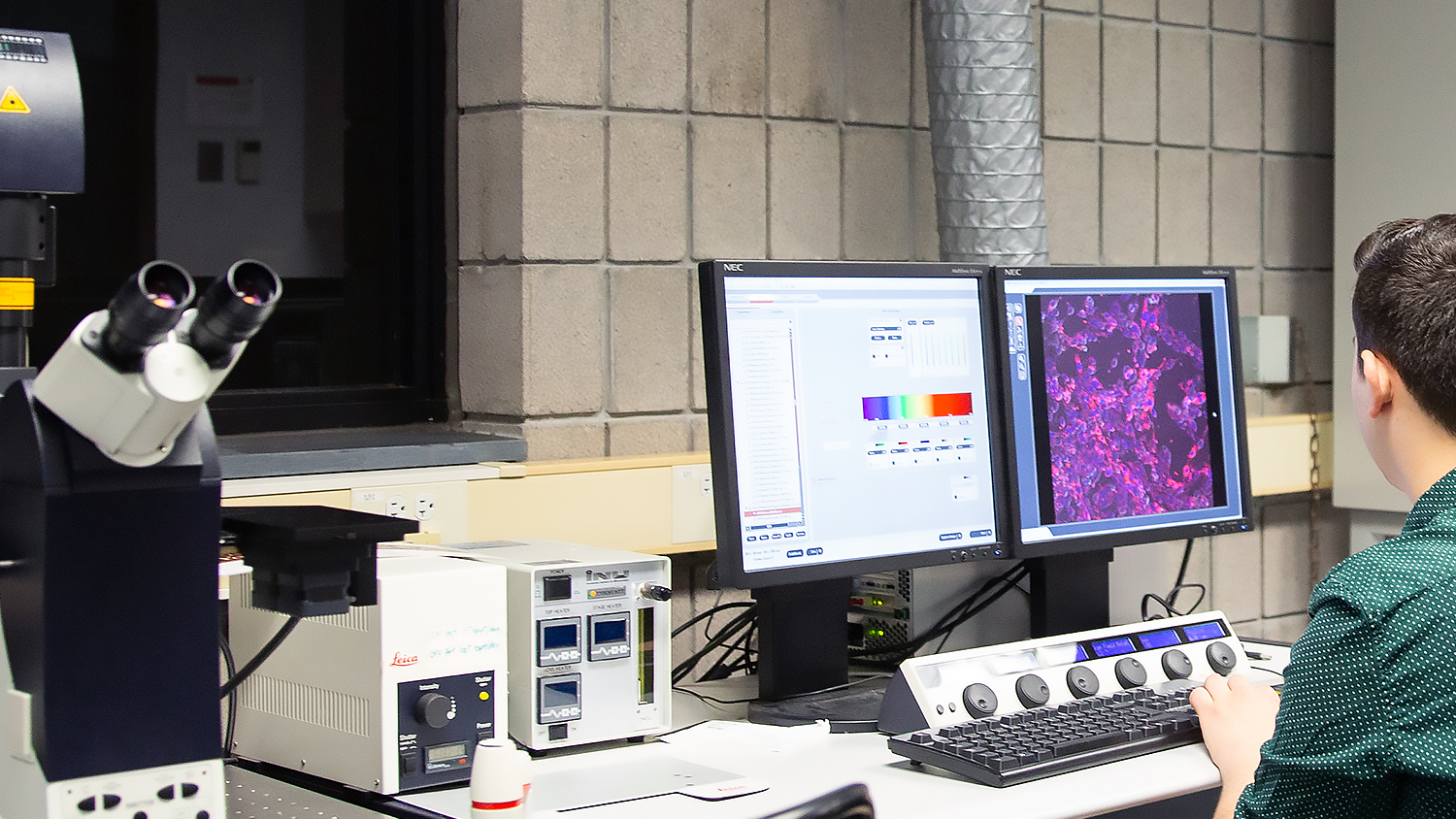
State-of-the-Art Genomics Lab: Access our Genomics Lab equipped with an Illumina MiSeq, allowing students to sequence and annotate whole genomes of diverse organisms, providing hands-on experience with cutting-edge technology in genomic analysis.
RIT’s bioinformatics master’s degree combines biotechnology, computer programming, and computational mathematics to prepare you to utilize and create technologies that discover, treat, and cure a range of medical illnesses. With a strong foundation in biotechnology, computer programming, computational mathematics, statistics, and database management, you will be well-prepared for academia and careers in the biotechnology, bioinformatics, pharmaceutical, and vaccine industries.
RIT’s Bioinformatics Master’s Degree
Bioinformatics is a field that has been developing over the last thirty years. It is a discipline that represents a marriage between biotechnology and computer technologies and has evolved through the convergence of advances in each of these fields. Today bioinformatics is a field that encompasses all aspects of the application of computer technologies to biological data. Computers are used to organize, link, analyze, and visualize complex sets of biological data to discover, treat, and cure a range of medical illnesses.
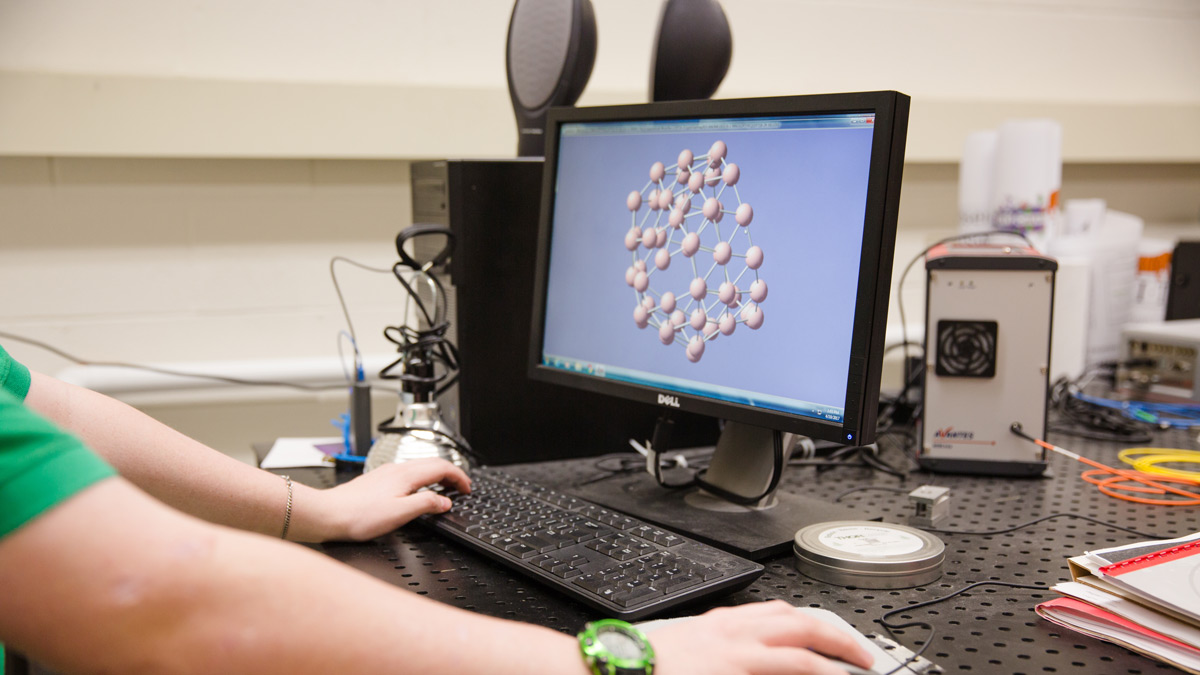
RIT’s bioinformatics master’s degree is focused on cutting-edge computational techniques, such as data mining, to understand biomedical data. In laboratory exercises and assignments, you will learn to sequence DNA and use computer programs to analyze DNA sequences and predict molecular models. You are also encouraged to pursue cooperative education opportunities to gain hands-on career experience in industry.
Current bioinformatics students have worked on projects including:
- Database development
- Cancer vaccine design
- Literature mining
- Molecular dynamics simulation
The program provides you with the capability to enter the bioinformatics workforce and become a leader in the field. The curriculum is designed to fulfill the needs of students with diverse educational and professional backgrounds. Individuals entering the program typically have degrees in biology, biotechnology, chemistry, statistics, computer science, information technology, or a related field. To prepare applicants from various backgrounds, the curriculum includes a comprehensive bridge program that includes courses in biology, mathematics, computer science, statistics, or other related fields. The program offers two tracks, one for students with backgrounds in the life sciences and one for those with backgrounds in the computational sciences.
Careers in Bioinformatics
With the advent of high-throughput technologies such as Next Generation Sequencing and proteomics, bioinformatics has become essential to the biological sciences in general. In the past, laboratories were able to manage and analyze their experimental data in spreadsheets. Many research labs now require the expertise of dedicated bioinformatics core centers or their own in-house bioinformaticists.
Graduates of the bioinformatics master’s program have entered such laboratories, both in industry and academia, as bioinformaticists. Some have also gone on to leverage their biotechnology experiences as wet lab experimentalists. The diversity of skills you will cultivate in the program give you access to a wide range of career choices.
The job market is rich with opportunities for those with graduate degree in bioinformatics, particularly when coupled with research as thesis work. This research provides exposure to real-world problems—and their solutions—not otherwise attainable in an academic setting.
Graduates of the bioinformatics master’s degree currently work senior analysts/programmers, associate systems analysts, bioinformaticist, bioinformatics analysts, bioinformatics engineers, computational biologists, and software engineers.
-
Affordable Now. Valuable for Life.
Earn your master’s degree without the full price tag. With Master Up you can receive a 30% tuition scholarship for an RIT master’s degree.
-
Join us for Fall 2026
Many programs accept applications on a rolling, space-available basis.
Careers and Cooperative Education
Typical Job Titles
| Researcher | Business Intelligence Developer | Computational Biologist |
| Bioinformatics Software Engineer | RD Bioinformatics and Laboratory Researcher | Innovation Consultant |
| Associate Software Engineer |
Cooperative Education
What makes an RIT science and math education exceptional? It’s the ability to complete science and math co-ops and gain real-world experience that sets you apart. Co-ops in the College of Science include cooperative education and internship experiences in industry and health care settings, as well as research in an academic, industry, or national lab. These are not only possible at RIT, but are passionately encouraged.
What makes an RIT education exceptional? It’s the ability to complete relevant, hands-on career experience. At the graduate level, and paired with an advanced degree, cooperative education and internships give you the unparalleled credentials that truly set you apart. Learn more about graduate co-op and how it provides you with the career experience employers look for in their next top hires.
Featured Work and Profiles
-
Transforming Genomic Data into Cancer Insight
RIT alum Spencer Richman ’20 uses computing and biology to build software that transforms genomic data into tools for cancer detection, helping bridge data and discovery in healthcare.
Read More about Transforming Genomic Data into Cancer Insight -
Dadgar Works to Make Medicine Personal
Sherry Dadgar launched PMCDx in 2020 to deliver advanced clinical genomic diagnostic testing to patients and their physicians. Today, PMCDx has more than 20 employees.
Read More about Dadgar Works to Make Medicine Personal -
Love Biology? Hate Formaldehyde? Try Bioinformatics.
Jeselle Clark realized there are biology career paths outside of medicine or ecology. Today she’s a bioinformatics software engineer working at Essex Management, LLC as a contractor for the National...
Read More about Love Biology? Hate Formaldehyde? Try Bioinformatics. -
Bioinformatics: The Intersection of Biology and Computer Science
Spencer Richman ‘20 switched majors when he discovered the bioinformatics program at RIT combined the two things he loved—computer science and biology.
Read More about Bioinformatics: The Intersection of Biology and Computer Science -
A Team Experience That Pays Off In More Ways Than One
Mary Pryor The Laboratory Support Team (or BioPrep) is a unique team that gets hands-on lab experience while helping the many teaching labs in the Thomas H. Gosnell School of Life Sciences at RIT.
Read More about A Team Experience That Pays Off In More Ways Than One
Curriculum for 2025-2026 for Bioinformatics MS
Current Students: See Curriculum Requirements
Students are also interested in
Admissions and Financial Aid
This program is available on-campus only.
| Offered | Admit Term(s) | Application Deadline | STEM Designated |
|---|---|---|---|
| Full‑time | Fall or Spring | Rolling | Yes |
| Part‑time | Fall or Spring | Rolling | No |
Full-time study is 9+ semester credit hours. Part-time study is 1‑8 semester credit hours. International students requiring a visa to study at the RIT Rochester campus must study full‑time.
Application Details
To be considered for admission to the Bioinformatics MS program, candidates must fulfill the following requirements:
- Complete an online graduate application.
- Submit copies of official transcript(s) (in English) of all previously completed undergraduate and graduate course work, including any transfer credit earned.
- Hold a baccalaureate degree (or US equivalent) from an accredited university or college in biology, biotechnology, biochemistry, chemistry, computer science, information technology, statistics, or a related discipline. A minimum cumulative GPA of 3.2 (or equivalent) is recommended.
- Submit a current resume or curriculum vitae.
- Submit a personal statement of educational objectives.
- Submit two letters of recommendation.
- Entrance exam requirements: GRE required for individuals with degrees from international universities
- Submit English language test scores (TOEFL, IELTS, PTE Academic, etc.), if required. Details are below.
English Language Test Scores
International applicants whose native language is not English must submit one of the following official English language test scores. Some international applicants may be considered for an English test requirement waiver.
Duolingo (DET): 120
IELTS: 6.5
LanguageCert Academic: 70
PTE Academic: 56
TOEFL: 79/4.5
International students below the minimum requirement may be considered for conditional admission. Deaf and hard-of-hearing test takers with significant hearing loss do not need to take the listening and speaking sections for the TOEFL and IELTS. Each program requires balanced sub-scores when determining an applicant’s need for additional English language courses.
How to Apply Start or Manage Your Application
Cost and Financial Aid
An RIT graduate degree is an investment with lifelong returns. Graduate tuition varies by degree, the number of credits taken per semester, and delivery method. View the general cost of attendance or estimate the cost of your graduate degree.
A combination of sources can help fund your graduate degree. Learn how to fund your degree
Accreditation
Research
Faculty in the College of Science receive research grant awards from organizations such as the National Science Foundation and the National Institutes of Health, which provide you with unique opportunities to conduct cutting-edge graduate-level thesis research. Using modern computational tools and approaches the bioinformatics faculty conduct research on a broad variety of topics including:
- molecular evolution
- ecological modeling
- cancer biology
- genomics
Learn more by exploring our life science research areas.
Related News
-
December 2, 2025

Students go Into the ROC to connect with their community
Through Into the ROC, a program that introduced nearly 8,500 RIT community members to the region since 2016, students have explored, volunteered, and built lasting community ties.
-
April 25, 2023

RIT graduate programs rank among best in nation in ‘U.S. News & World Report’ 2023 survey
RIT graduate degree programs are among the best in the nation, according to the U.S. News & World Report annual statistical survey of graduate programs.
-
December 2, 2022

Study by RIT scientists indicates SARS-CoV-2 variants are still transmissible between species
Scientists believe bats first transmitted SARS-CoV-2 to humans in December 2019, and while the virus has since evolved into several variants such as delta and omicron, a new study by scientists at RIT indicates the virus is still highly transmissible between mammals.
Contact
- Lindsay Lewis
- Senior Assistant Director
- Graduate Admissions
- Enrollment Management
- 585‑475‑5532
- lslges@rit.edu
- Crista Wadsworth
- Associate Professor, Biology
- Thomas H. Gosnell School of Life Sciences
- College of Science
- 585‑475‑7961
- cbwsbi@rit.edu
Thomas H. Gosnell School of Life Sciences